Reconstituting the Bacterial Cytoskeleton: A Protocol for FtsZ and Actin Dynamics in GUVs
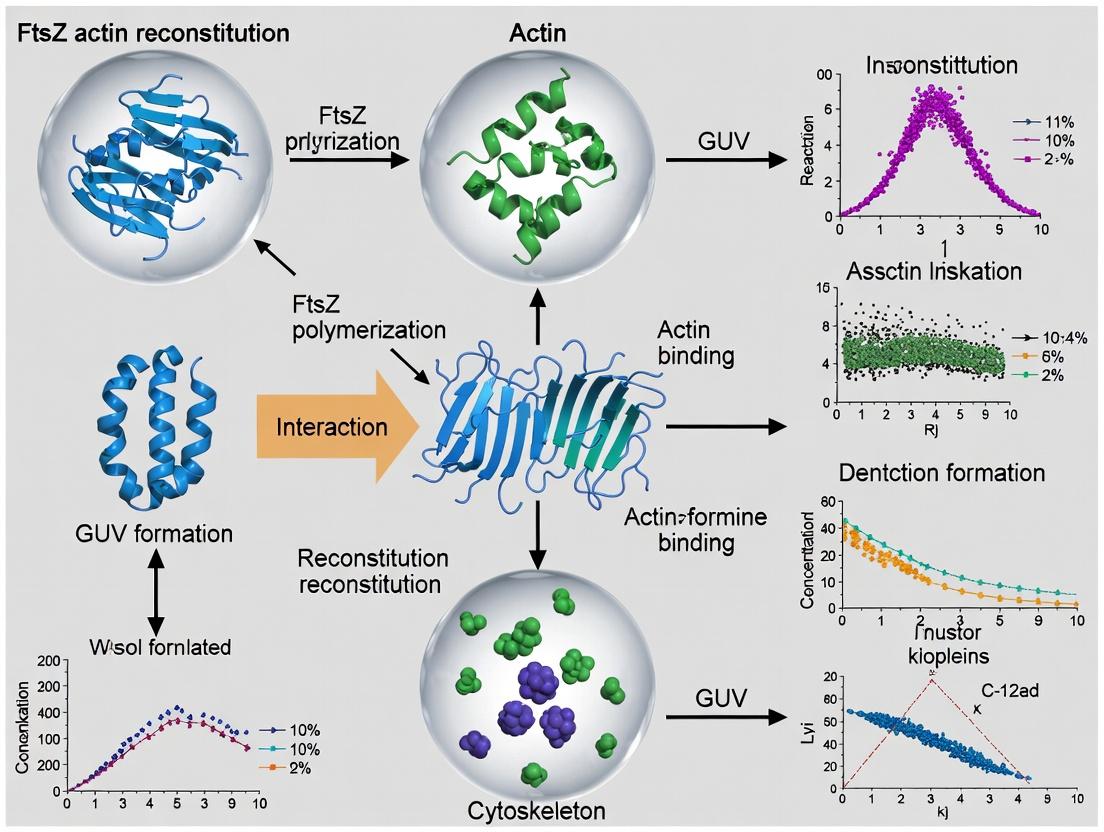
This article provides a comprehensive guide for researchers aiming to reconstitute key bacterial cytoskeletal proteins, FtsZ and actin-like MreB, inside Giant Unilamellar Vesicles (GUVs).
Reconstituting the Bacterial Cytoskeleton: A Protocol for FtsZ and Actin Dynamics in GUVs
Abstract
This article provides a comprehensive guide for researchers aiming to reconstitute key bacterial cytoskeletal proteins, FtsZ and actin-like MreB, inside Giant Unilamellar Vesicles (GUVs). We cover the foundational biology of these proteins, detail a robust methodological pipeline for encapsulation and visualization, address common experimental pitfalls with optimization strategies, and discuss validation techniques and comparative analyses with other model systems. Targeted at scientists in biophysics, synthetic biology, and antimicrobial drug discovery, this guide synthesizes current best practices to enable the study of division and morphogenesis machinery in a controlled, cell-like environment.
The Bacterial Cytoskeleton Unveiled: Understanding FtsZ, Actin, and GUV Fundamentals
This document provides essential context and methodologies for research investigating the reconstitution of bacterial cell division and morphogenesis in Giant Unilamellar Vesicles (GUVs). The work is framed within a broader thesis aiming to construct a minimal synthetic cell, focusing on the cytoskeletal proteins FtsZ (tubulin homolog) and MreB (actin homolog). FtsZ is the primary driver of bacterial cytokinesis, assembling into the Z-ring that constricts the membrane. Actin-like proteins, such as MreB, are crucial for defining and maintaining rod-shaped cell morphology and coordinating peptidoglycan synthesis. Reconstituting these systems in GUVs allows for the dissection of their minimal requirements, interactions, and potential for targeted disruption in drug development.
Recent Findings (Live Search Summary): Recent advances (2023-2024) highlight the use of engineered FtsZ variants with enhanced membrane binding (e.g., via FtsA-mimicking peptides or direct lipid anchors) to achieve constriction in GUVs. For morphogenesis, co-reconstitution of MreB filaments with membrane-linked peptidoglycan synthases in lipid tubules has shown promise in creating defined shapes. Key quantitative parameters from recent literature are summarized below.
Table 1: Key Quantitative Parameters for FtsZ & MreB Reconstitution
| Parameter | FtsZ System (Typical Range) | MreB System (Typical Range) | Significance |
|---|---|---|---|
| Protein Concentration | 2 - 10 µM (monomer) | 1 - 5 µM (monomer) | Optimal for filament assembly without nonspecific aggregation. |
| GUV Lipid Composition | DOPC + 5-20% charged lipid (e.g., DOPG, CL) + 1-2% FtsZ anchor (e.g., DOGS-NTA-Ni²⁺ for His-tagged FtsZ) | DOPC + MreB anchor (e.g., lipid-linked undecaprenol analogs) + Rod-shaped template (optional) | Provides membrane stability, protein binding sites, and geometric cues. |
| Critical Assembly Concentration (CAC) | ~0.5 - 1.2 µM (varies with anchors, GTP, crowding agents) | ~0.2 - 0.8 µM (varies with ATP, Mg²⁺, membrane linkage) | Minimum concentration for productive filament assembly on membranes. |
| Constriction/Deformation Rate | 1 - 10 nm/s (in GUVs with anchored FtsZ) | Tubule elongation: 0.05 - 0.5 µm/min (in vitro assays) | Measure of cytoskeletal force generation and dynamics. |
| Optimal Ionic Conditions | 50-100 mM KCl, 5-10 mM MgCl₂, 1-2 mM GTP, pH 7.0-7.5 | 50-150 mM KCl, 2-5 mM MgCl₂, 1-5 mM ATP, pH 7.2-7.7 | Supports polymerization, nucleotide hydrolysis, and membrane interaction. |
| Key Crowding Agent | 2-4% w/v PEG (MW 8000) or Ficoll 400 | 1-3% w/v Methylcellulose or BSA | Mimics cytoplasmic crowding, stabilizing filaments and bundling. |
Experimental Protocols
Protocol 2.1: Reconstitution of Membrane-Anchored FtsZ Constriction in GUVs
Objective: To assemble GTP-dependent FtsZ filaments on the inner leaflet of GUVs and observe membrane constriction.
Materials: See "The Scientist's Toolkit" (Section 4). Buffer A (Interior/Assembly Buffer): 50 mM HEPES, 100 mM KCl, 10 mM MgCl₂, 1 mM GTP, 2% PEG 8000, pH 7.2. Buffer B (Exterior/Osmotic Support Buffer): 50 mM HEPES, 100 mM KCl, 200 mM sucrose, pH 7.2.
Method:
- GUV Preparation (Electroformation): Prepare a lipid mixture in chloroform: DOPC (78 mol%), DOPG (20 mol%), DOGS-NTA-Ni (2 mol%). Dry 10 µl of lipid solution (5 mg/ml) on indium tin oxide (ITO)-coated glass slides. Assemble a chamber, fill with 300 mOsm sucrose solution, and apply an AC field (1 V, 10 Hz, 2 hours) at 60°C. Harvest GUVs gently.
- FtsZ Protein Preparation: Express and purify His-tagged FtsZ (or FtsZ fused to an amphipathic helix anchor) using standard Ni-NTA chromatography. Dialyze into Buffer A without GTP and concentrate to 20 µM. Keep on ice.
- Membrane Binding & Constriction Assay: a. In a perfusion chamber, mix 50 µl of harvested GUVs with 50 µl of Buffer B containing 10 mM NiCl₂ to charge the NTA lipids. Incubate for 5 min. b. Gently wash with 200 µl Buffer B to remove excess Ni²⁺. c. Dilute the FtsZ stock 1:1 in Buffer A with GTP to achieve a final working concentration of 5 µM FtsZ, 1 mM GTP. d. Perfuse 100 µl of the FtsZ/GTP solution into the chamber. e. Immediately transfer to a TIRF or confocal microscope stage pre-warmed to 30°C. f. Acquire time-lapse images (30-60 sec intervals for 30-60 min).
- Analysis: Measure GUV diameter over time using image analysis software (e.g., Fiji). Constriction is defined as a ≥20% reduction in diameter.
Protocol 2.2: Reconstitution of MreB-Directed Morphogenesis on Membrane Templates
Objective: To assemble membrane-bound MreB filaments and observe their role in shaping lipid structures.
Materials: See "The Scientist's Toolkit" (Section 4). MreB Buffer (MB): 25 mM HEPES, 150 mM KCl, 5 mM MgCl₂, 5 mM ATP, 1% Methylcellulose (viscosity enhancer), pH 7.5.
Method:
- Template Preparation (Rod-shaped SLBs): Create rod-shaped silica or PDMS microchambers (1-2 µm width). Form a supported lipid bilayer (SLB) inside by vesicle fusion using a lipid mix containing 95% DOPC and 5% lipid capable of binding MreB (e.g., biotinylated PE for streptavidin-linked MreB, or specific glycolipids).
- MreB Protein Preparation: Express and purify MreB (often with a membrane-targeting tag like an N-terminal amphipathic helix or a biotinylation tag for streptavidin linkage). Clarify by high-speed centrifugation (150,000 x g, 30 min) before use to remove aggregates.
- Morphogenesis Assay: a. Dilute purified MreB in MB to a final concentration of 3 µM. b. Introduce the MreB solution into the microchamber containing the rod-shaped SLB template. c. Incubate at 30°C for 15-30 minutes without flow. d. Image using TIRF microscopy to visualize filament alignment along the long axis of the template. e. For dynamic studies, perform fluorescence recovery after photobleaching (FRAP) on MreB filaments to assess turnover.
- Analysis: Quantify filament orientation relative to the template's long axis. Measure the persistence length of filaments on the membrane.
Visualizations
FtsZ Assembly and Constriction Pathway
GUV Reconstitution Workflow
The Scientist's Toolkit
Table 2: Essential Research Reagent Solutions
| Item | Function in Reconstitution | Example Product/Specification |
|---|---|---|
| 1,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC) | Major uncharged lipid for forming stable, fluid GUV membranes. | Avanti Polar Lipids, 850375C. >99% purity. |
| 1,2-dioleoyl-sn-glycero-3-phospho-(1'-rac-glycerol) (DOPG) | Negatively charged lipid, essential for recruiting cationic proteins and mimicking bacterial membrane charge. | Avanti Polar Lipids, 840475C. |
| DOGS-NTA-Ni²⁺ Lipid | Synthetic lipid for anchoring His-tagged proteins (e.g., FtsZ) to the GUV membrane via Ni²⁺ coordination. | Avanti Polar Lipids, 790404C. |
| PEG 8000 | Macromolecular crowding agent. Mimics the excluded volume effect of the cytoplasm, promoting FtsZ filament bundling and Z-ring maturation. | Sigma-Aldrich, 89510. |
| Methylcellulose (4000 cP) | Viscosity enhancer and crowding agent for MreB assays. Reduces filament fragmentation and supports network formation on membranes. | Sigma-Aldrich, M0512. |
| GTP (Guanosine Triphosphate) | Hydrolyzable nucleotide fuel for FtsZ polymerization and dynamic treadmilling. Critical for force generation. | Jena Bioscience, NU-1012. High-purity, lithium salt. |
| ATP (Adenosine Triphosphate) | Hydrolyzable nucleotide for MreB polymerization and dynamic assembly. Required for morphogenetic activity. | Roche, 10127531001. |
| His-tagged FtsZ Protein | Core cytoskeletal protein. Recombinant expression (often from E. coli) with a hexahistidine tag for purification and optional membrane anchoring. | Purified via Ni-NTA, >95% purity, stored in -80°C. |
| MreB Protein (with membrane tag) | Actin-like protein for morphogenesis. Requires engineering (e.g., N-terminal amphipathic helix, biotin tag) for stable membrane association. | Purified via affinity chromatography, clarified by ultracentrifugation. |
| Sucrose/Glucose Osmotic Buffer Pair | Creates an osmotic gradient to stabilize GUVs during microscopy. Sucrose inside, glucose outside matches refractive index for better imaging. | Osmolarity verified with a micro-osmometer (~300 mOsm). |
Why GUVs? The Advantages of a Minimal Synthetic Cell Platform
Within the broader thesis investigating the reconstitution of FtsZ and actin cytoskeletal networks for synthetic cell division, Giant Unilamellar Vesicles (GUVs) serve as the foundational platform. This application note details why GUVs are the optimal chassis for this research, providing a detailed comparison of their advantages and robust protocols for their use in cytoskeletal reconstitution studies.
Key Advantages of GUVs: A Quantitative Comparison
GUVs offer distinct benefits over other membrane models (like SUVs, LUVs, or supported bilayers) and living cells for minimal synthetic cell research. The quantitative and qualitative data below summarize these advantages.
Table 1: Comparative Analysis of Membrane Model Systems
| Feature | GUVs | SUVs/LUVs | Supported Lipid Bilayers (SLBs) | Living Cells (E. coli/HeLa) |
|---|---|---|---|---|
| Size Range (Diameter) | 1 - 100 µm | 0.02 - 0.1 µm | N/A (2D sheet) | 1 - 100 µm |
| Compartmentalization | Yes, enclosed 3D lumen | Yes, but tiny lumen | No (2D) | Yes, complex |
| Membrane Curvature | Tunable (low) | Very high | Flat | Complex & dynamic |
| Asymmetric Leaflet Formation | Possible (advanced techniques) | Difficult | Possible | Intrinsic |
| Cytoskeletal Protein Reconstitution | Ideal for FtsZ/actin rings | Not feasible | Limited to 2D dynamics | Endogenous background |
| Microscopy Compatibility | Excellent for phase/confocal | Poor (too small) | Good (TIRF) | Good but crowded |
| Content Encapsulation Efficiency | Moderate (1-10%) | High | N/A | N/A |
| Throughput for Analysis | Moderate (100s/experiment) | High (bulk) | High | High |
| System Complexity | Minimal, defined | Minimal | Minimal | Extremely High |
Table 2: Quantitative Metrics for GUVs in Synthetic Cell Research
| Metric | Typical Value/Result | Implication for FtsZ/Actin Studies |
|---|---|---|
| Membrane Bending Rigidity | ~20 kBT (for DOPC) | Similar to natural membranes; affects filament deformation. |
| Successful FtsZ Ring Formation* | 30-70% of GUVs (literature range) | Demonstrates feasibility of cytokinesis mimicry. |
| Actin Cortex Reconstitution* | Achieved on ~60% of GUVs | Enables study of membrane-cortex mechanical coupling. |
| Electroformation Yield | 107 - 108 GUVs/mL | Sufficient for bulk biochemical assays and microscopy. |
| Stability at Room Temp | 24 - 72 hours | Allows for extended time-lapse experiments. |
| Permeability Control | Tunable via lipid choice/cholesterol | Enables triggered activation of internal reactions. |
*Dependent on protein quality, lipid composition, and encapsulation method.
Core Protocols for GUV-based Reconstitution
Protocol 3.1: Electroformation of GUVs for Cytoskeletal Studies
Objective: Produce monodisperse, unilamellar GUVs in an isotonic sucrose solution for subsequent protein encapsulation or external protein addition.
Research Reagent Solutions:
- Lipid Stock Solutions: 10 mg/mL DOPC, DOPS, DOPE, and cholesterol in chloroform. Function: Define membrane fluidity, charge, and stability.
- Electroformation Buffer (Internal): 200 mM sucrose. Function: Creates osmolarity for later exchange; compatible with microscopy.
- Observation/Glucose Buffer (External): 200 mM glucose, 1-10 mM MgCl2, 50 mM HEPES (pH 7.4). Function: Creates density difference for vesicle settling; Mg2+ aids protein-membrane binding.
Methodology:
- Indium Tin Oxide (ITO) Slide Preparation: Clean ITO slides sequentially with ethanol, acetone, and Milli-Q water. Dry under nitrogen.
- Lipid Deposition: Mix lipid stocks to desired composition (e.g., 70% DOPC, 15% DOPS, 15% DOPE) in a glass vial. Using a Hamilton syringe, spread 10-20 µL of the lipid mix evenly onto the conductive side of an ITO slide. Dry under vacuum for 1 hour to remove all organic solvent.
- Chamber Assembly: Assemble a Teflon spacer (2 mm thick) between the lipid-coated slide and a clean ITO slide. Secure with binder clips.
- Vesicle Growth: Fill the chamber with pre-warmed (37°C) electroformation buffer (sucrose). Connect electrodes to a function generator. Apply a low-frequency AC field (10 Hz, 1.1 V) for 1 hour at 37°C, then switch to 2 Hz for 1-2 hours.
- Harvesting: Carefully collect the GUV-containing sucrose solution using a cut pipette tip. Transfer to an Eppendorf tube.
- Osmotic Stabilization: Gently layer an equal volume of observation buffer (glucose) beneath the GUV suspension. Centrifuge at 300 x g for 5 minutes. The GUVs will settle at the interface/bottom. Carefully collect for experiments.
Protocol 3.2: Active Cytoskeletal Protein Reconstitution on GUVs
Objective: Assemble functional FtsZ or actin filaments on the inner or outer leaflet of GUVs to study division machinery.
Research Reagent Solutions:
- 10X Reconstitution Buffer: 500 mM KCl, 50 mM MgCl2, 10 mM ATP/GTP, 200 mM HEPES (pH 7.2). Function: Provides optimal ionic conditions for FtsZ/actin polymerization.
- Fluorescently Labeled FtsZ/Actin: Purified protein labeled with Alexa Fluor 488/568. Function: Enables real-time visualization of filament dynamics via fluorescence microscopy.
- Membrane-Tethering Agent: Biotinylated lipids (e.g., DOPE-biotin) + NeutrAvidin, or His-tagged proteins + Ni-NTA lipids. Function: Anchors cytoskeletal filaments to the membrane to generate force.
Methodology for External Cortex Assembly (e.g., FtsZ):
- Prepare GUVs in observation buffer as per Protocol 3.1.
- In a separate tube, mix purified FtsZ (2-5 µM final) in 1X Reconstitution Buffer (with GTP).
- Place 20 µL of the GUV suspension on a clean coverslip. Add 20 µL of the FtsZ mix. Mix gently by pipetting.
- Immediately image using TIRF or confocal microscopy at 25-30°C. Monitor for the formation of continuous, contracting filament rings at the GUV equator.
Methodology for Encapsulation & Internal Assembly (e.g., Actin):
- Hydration during Electroformation: Use electroformation buffer containing 1-5 µM actin (monomers) and 1X Reconstitution Buffer (with ATP).
- Perform electroformation as in Protocol 3.1. This encapsulates actin monomers inside the GUVs.
- Triggered Polymerization: After harvesting, introduce a calcium chelator (EGTA) or increase ionic strength in the external buffer to trigger actin polymerization inside the GUV lumen, potentially forming a cortical network.
Visualization of Experimental Workflows
Title: GUV Synthesis & Protein Reconstitution Workflow
Title: Minimal FtsZ Ring Assembly Pathway on GUVs
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for FtsZ/Actin-GUV Reconstitution
| Reagent/Category | Specific Example(s) | Function & Rationale |
|---|---|---|
| Lipids | DOPC, DOPS, DOPE, Cholesterol, DOPE-biotin, Ni-NTA-DGS | DOPC: Provides neutral, fluid bilayer backbone. DOPS: Introduces negative charge for FtsZ/membrane interaction. DOPE: Promotes non-lamellar phases, mimicking bacterial membrane stress. Cholesterol: Modifies membrane rigidity and permeability. Functionalized Lipids: Enable specific protein tethering. |
| Polymerization Buffers | HEPES (pH 7.2-7.4), KCl, MgCl2, GTP, ATP, CaCl2/EGTA | Mg2+: Essential cation for FtsZ/actin polymerization and membrane binding. GTP/ATP: Hydrolyzable fuel for cytoskeletal dynamics. Ca2+/EGTA: Used as a switch to trigger actin polymerization inside GUVs. |
| Purified Proteins | FtsZ (WT/mutants), Actin (G-/F-), Nucleating Factors (FtsA, formins), Cross-linkers | Core structural components. Mutants allow probing mechanics. Nucleators control assembly location and kinetics. Cross-linkers (e.g., α-actinin) stabilize networks. |
| Fluorescent Labels | Alexa Fluor 488/568/647 maleimide, ATTO dyes | Site-specific labeling of proteins for dynamic, quantitative fluorescence microscopy without disrupting function. |
| Microscopy Substrates | Passivated glass coverslips, PEG-silane, BSA treatment | Prevents GUV and protein adhesion to the glass surface, ensuring free-floating vesicles and minimizing artifacts. |
| GUV Production Equipment | ITO-coated glass slides, function generator, temperature chamber | Standard setup for reliable, high-yield GUV formation via the gentle hydration electroformation method. |
Application Notes: Framework for FtsZ Actin Reconstitution in GUVs
Reconstituting the bacterial cytoskeletal protein FtsZ within Giant Unilamellar Vesicles (GUVs) serves as a minimal system to study the physical principles of bacterial cell division. This platform is crucial for fundamental biophysical research and for screening compounds that modulate FtsZ polymerization—a promising antibacterial target. The success of such bottom-up synthetic biology approaches hinges on two pillars: the production of high-purity, functional protein and the formation of suitable biomimetic lipid compartments.
Key Challenges & Considerations:
- Protein Purity & Activity: Contaminants like proteases or nucleotidases can degrade FtsZ or its GTP fuel, leading to failed polymerization. Tagged purification strategies are essential.
- Lipid Composition: The membrane must be compatible with FtsZ recruitment. This often requires incorporating lipids with anionic headgroups (e.g., PG, CL) to interact with the protein's positively charged C-terminal tail. Membrane fluidity and curvature are also critical parameters.
- Internal Solution & Buffering: The reconstitution buffer must support both protein activity and membrane integrity, often requiring optimized salt, pH, and crowding agents.
Detailed Protocols
Protocol: His-Tagged FtsZ Protein Purification (IMAC)
This protocol details the purification of recombinant FtsZ from E. coli using Immobilized Metal Affinity Chromatography (IMAC).
Materials:
- E. coli BL21(DE3) cells expressing His₆-FtsZ
- Lysis Buffer: 50 mM Tris-HCl pH 7.9, 500 mM NaCl, 10 mM imidazole, 1 mM PMSF, 1 mg/mL lysozyme.
- Wash Buffer: 50 mM Tris-HCl pH 7.9, 500 mM NaCl, 30 mM imidazole.
- Elution Buffer: 50 mM Tris-HCl pH 7.9, 500 mM NaCl, 300 mM imidazole.
- Storage/Dialysis Buffer: 50 mM HEPES-KOH pH 7.2, 200 mM KCl, 1 mM MgCl₂, 1 mM DTT.
- Ni-NTA Agarose Resin
- FPLC or Gravity Column System
Method:
- Cell Lysis: Resuspend cell pellet in Lysis Buffer. Incubate on ice for 30 min. Lyse by sonication (5x 30s pulses, 70% amplitude). Clarify lysate by centrifugation at 20,000 x g for 45 min at 4°C.
- Column Preparation: Equilibrate 2 mL Ni-NTA resin with 10 column volumes (CV) of Lysis Buffer.
- Binding: Incubate clarified lysate with equilibrated resin for 1 hour at 4°C with gentle mixing.
- Washing: Load resin into a column. Wash with 10 CV of Wash Buffer until A₂₈₀ baseline stabilizes.
- Elution: Elute bound protein with 5 CV of Elution Buffer. Collect 1 mL fractions.
- Buffer Exchange & Storage: Pool fractions containing FtsZ (confirmed by SDS-PAGE). Dialyze overnight into Storage Buffer at 4°C. Concentrate using a centrifugal filter (MWCO 30 kDa). Aliquot, snap-freeze in liquid N₂, and store at -80°C. Determine concentration by A₂₈₀ (ε ≈ 43,860 M⁻¹cm⁻¹).
Protocol: Lipid Selection and GUV Formation via Electroformation
This protocol describes the formation of GUVs with defined lipid compositions suitable for FtsZ reconstitution.
Materials:
- Lipids: DOPC, DOPE, DOPS, Cardiolipin (CL) in chloroform stock solutions.
- Electroformation Buffer: 200 mM sucrose, 10 mM HEPES pH 7.2.
- External (Glucose) Buffer: 200 mM glucose, 10 mM HEPES pH 7.2.
- Indium Tin Oxide (ITO)-coated glass slides.
- Electroformation chamber.
- Function generator.
Method:
- Lipid Film Preparation: Mix lipid chloroform stocks in a glass vial to achieve desired molar ratio (e.g., DOPC:DOPE:DOPS:CL = 60:20:15:5). Dry under a stream of N₂ gas to form a thin film. Desiccate under vacuum for >2 hours.
- Slide Assembly & Hydration: Rehydrate the lipid film with ~20 µL of Electroformation Buffer (sucrose) to create a lipid slurry. Spread onto one ITO slide. Assemble the chamber with a second ITO slide using a 1-2 mm spacer. Fill chamber with sucrose buffer.
- Electroformation: Connect slides to a function generator. Apply an AC field: 1.1 V, 10 Hz, for 90-120 minutes at a temperature above the lipid phase transition (e.g., 28°C for DOPC mixtures).
- GUV Harvesting: Gently flush the chamber with ~1 mL of External (Glucose) Buffer. The density difference (sucrose inside, glucose outside) helps settle GUVs, improving visualization. Harvest GUVs using a wide-bore pipette tip.
Table 1: Common Lipid Compositions for FtsZ-Reconstitution GUVs
| Lipid Component | Abbreviation | Typical Mol% Range | Functional Role |
|---|---|---|---|
| 1,2-dioleoyl-sn-glycero-3-phosphocholine | DOPC | 50 - 70 % | Neutral, fluid bilayer matrix |
| 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine | DOPE | 15 - 30 % | Conical shape, promotes curvature |
| 1,2-dioleoyl-sn-glycero-3-phospho-L-serine | DOPS | 10 - 20 % | Anionic charge for FtsZ recruitment |
| Heart Extract Cardiolipin | CL | 0 - 10 % | High negative charge, membrane microdomain organization |
Table 2: Critical Parameters for FtsZ Polymerization in Vitro
| Parameter | Optimal Range | Impact on Activity |
|---|---|---|
| GTP Concentration | 1 - 5 mM | Polymerization fuel; higher [GTP] increases polymer turnover. |
| Mg²⁺ Concentration | 5 - 10 mM | Essential cofactor for GTP binding and polymerization. |
| KCl Concentration | 100 - 200 mM | Modulates polymerization kinetics and bundle morphology. |
| pH | 6.5 - 7.5 | Affects FtsZ protonation state and polymer stability. |
| Macromolecular Crowder (PEG) | 2 - 4 % (w/v) | Mimics cellular crowding, promotes bundling. |
Diagrams
Title: Workflow for FtsZ Reconstitution in GUVs
Title: FtsZ Polymerization & Membrane Interaction Pathway
The Scientist's Toolkit
Table 3: Key Reagent Solutions for FtsZ-GUV Reconstitution
| Reagent / Material | Function & Importance | Example Product / Note |
|---|---|---|
| HEPES-KOH Buffer (pH 7.2-7.5) | Standard physiological pH buffer for FtsZ activity and membrane stability. Prevents pH drift. | Prepare 1M stock, filter sterilize. |
| High-Purity GTP | Essential nucleotide fuel for FtsZ polymerization. Requires >95% purity to avoid inhibition. | Sigma-Aldrich, Roche, in 100 mM aliquots at -80°C. |
| DTT or TCEP | Reducing agent. Maintains FtsZ cysteines in reduced state, critical for function. | Use fresh 1M DTT or 0.5M TCEP stocks. |
| Sucrose & Glucose (Osmotic Pair) | Creates density difference for GUV handling (sucrose inside, glucose outside). Osmolarity must be matched (~200 mOsm). | Use high-purity, prepare in buffer. |
| DOPC, DOPS, CL Lipids | Building blocks for anionic, fluid GUVs. High purity (>99%) ensures reproducible electroformation. | Avanti Polar Lipids, in chloroform under N₂ at -20°C. |
| Ni-NTA Agarose | IMAC resin for efficient His-tagged FtsZ purification. High binding capacity is crucial for yield. | Qiagen, Cytiva. |
| Protease Inhibitor Cocktail | Protects FtsZ from degradation during purification from E. coli. | EDTA-free recommended if using IMAC. |
| PEG (e.g., PEG 8000) | Macromolecular crowder. Mimics cellular environment, promotes FtsZ filament bundling. | Add to reaction buffer from 50% stock. |
This document provides application notes and protocols for investigating the in vitro reconstitution of the prokaryotic tubulin homolog FtsZ within Giant Unilamellar Vesicles (GUVs). This work is a core component of a broader thesis aiming to reconstitute minimal divisome machinery to understand the fundamental principles of bacterial cytokinesis and to establish a platform for screening potential antimicrobial agents that target this essential process.
Expected Polymerization Dynamics: Quantitative Parameters
The polymerization of FtsZ follows a nucleation-elongation model, sensitive to nucleotide (GTP), ionic conditions, and macromolecular crowding. Key quantitative parameters are summarized below.
Table 1: Core Parameters of FtsZ Polymerization Dynamics In Vitro
| Parameter | Typical Range in vitro (Buffer) | Influence on Polymerization | Notes for GUV Reconstitution |
|---|---|---|---|
| Critical Concentration (Cc) | 0.5 - 2.0 µM (with GTP/Mg²⁺) | Below Cc, no polymerization; above Cc, polymer mass increases linearly. | Intragvesicular concentration must exceed Cc. Achieved via active loading or de novo expression. |
| Nucleotide (GTP) Kinetics | KD ~ 0.1 - 0.5 µM; Hydrolysis rate ~ 4-6 GTP/min/FtsZ | GTP binding promotes assembly; hydrolysis promotes disassembly and treadmilling. | GTP regeneration systems often required to sustain dynamics. |
| Monomer Association Rate (kon) | ~ 5-10 µM⁻¹s⁻¹ | Governs elongation speed. | Affected by viscosity inside GUVs. |
| Monomer Dissociation Rate (koff) | ~ 2-8 s⁻¹ (GTP-bound); higher for GDP-bound. | Governs shrinkage/turnover. | Key parameter for treadmilling velocity. |
| Treadmilling Velocity | 10 - 30 nm/s in vitro; up to 50 nm/s in vivo. | Measure of net flux of subunits through filament. | Primary readout for functional reconstitution; measurable via TIRF on GUVs. |
| Cation Dependence (Mg²⁺) | Optimal ~ 2-10 mM | Essential for GTP binding/hydrolysis and polymer stability. | Must be present in internal GUV buffer. |
| Crowding Agent Effect (e.g., PEG) | 1-5% w/v can lower Cc 5-10 fold. | Mimics cytoplasmic crowdedness, promotes bundling. | Crucial for forming cohesive Z-rings versus scattered filaments. |
Expected Membrane Interactions: Anchoring Mechanisms
FtsZ must be linked to the membrane to generate constrictive force. Two primary reconstitution strategies are employed.
Table 2: Membrane Anchoring Strategies for FtsZ on GUVs
| Anchoring System | Components | Mode of Interaction | Key Experimental Considerations |
|---|---|---|---|
| Natural Protein Anchor (FtsA/FtsZ-CT) | FtsZ, FtsA (or ZipA), lipid with native headgroup (e.g., POPC/POPE). | FtsA binds membrane lipids (via amphipathic helix) and FtsZ C-terminal peptide (CTC). | Requires co-reconstitution of multiple proteins; more physiologically relevant. |
| Synthetic Lipid Anchor (e.g., DOGS-NTA-Ni²⁺) | His-tagged FtsZ, Ni²⁺-chelating lipid (e.g., 18:1 DGS-NTA(Ni)) doped into GUV membrane. | High-affinity coordination between Ni²⁺ and polyhistidine tag. | Simple, tunable (via lipid molar %), strong linkage but non-physiological. |
| Amphipathic Helix Fusion (e.g., FtsZ-MTS) | FtsZ fused to a membrane-targeting sequence (MTS) like MinD's MTS. | MTS inserts into lipid bilayer. | Direct linkage; may affect FtsZ polymerization if fusion site is suboptimal. |
| Cholesterol-Based Anchor (for lipid rafts) | His-tagged FtsZ, cholesterol-based NTA-lipid. | Targets anchors to liquid-ordered domains. | Useful for studying spatial organization in heterogeneous membranes. |
Detailed Experimental Protocols
Protocol 1: GUV Formation via Electroformation for FtsZ Reconstitution
Aim: To produce GUVs with a defined lipid composition suitable for protein reconstitution. Materials: Lipid stock solutions (e.g., POPC, POPG, DGS-NTA(Ni) at 1 mg/mL in chloroform), Pt electrodes, electroformation chamber, AC/DC power supply, sucrose/glucose solutions.
- Clean Electrodes: Sonicate Pt wires in ethanol, then water. Dry under N₂.
- Lipid Deposition: Spread 20 µL of lipid mix (e.g., 97% POPC, 3% DGS-NTA(Ni)) on each Pt electrode. Dry under vacuum for 1 hr to remove all solvent.
- Hydration: Assemble chamber with 500 µL of internal solution (e.g., 200 mM sucrose, 50 mM Tris-HCl pH 7.5, 50 mM KCl, 5-10 mM MgCl₂). Ensure electrodes are covered.
- Electroformation: Apply low-frequency AC field (10 Hz, 1.1 V) for 1 hr at 60°C (above lipid Tm), then 1.9 V for 30 min. Optionally, apply a 1 Hz frequency for final 5 min to detach GUVs.
- Harvesting: Carefully collect GUV solution from chamber. Keep on ice.
- Sedimentation: Layer GUV/sucrose solution under an equal volume of iso-osmotic glucose buffer in observation chamber. GUVs will settle to the interface.
Protocol 2: Reconstitution of His-FtsZ on NTA-GUVs and Treadmilling Assay
Aim: To anchor FtsZ polymers to GUVs and quantify their treadmilling dynamics. Materials: GUVs (with NTA-lipids) from Protocol 1, purified His₁₀-FtsZ, oxygen-scavenging system (0.5% glucose, 50 µg/mL glucose oxidase, 10 µg/mL catalase), GTP (1 mM), TIRF microscope.
- Pre-treatment of Observation Chamber: Passivate glass-bottom chamber with 1% BSA for 10 min, rinse.
- Sample Assembly in Chamber: Mix in order:
- 40 µL of glucose-based imaging buffer (with oxygen scavengers).
- 5 µL of settled GUVs (from Protocol 1, step 6).
- 5 µL of 10X FtsZ mix (in internal buffer): Final chamber concentration: 2 µM FtsZ, 1 mM GTP, 5 mM MgCl₂.
- Gently mix by pipetting.
- Incubation & Anchoring: Incubate for 5-10 min at 25°C to allow FtsZ polymerization and binding to NTA-lipids.
- TIRF Imaging: Image using a 488 nm laser for GFP-FtsZ or using suitable dye. Set acquisition to 1-2 frames/sec for 5-10 min.
- Treadmilling Analysis: Use kymograph analysis (Fiji/ImageJ) along the GUV circumference. The slope of fluorescent speckle movement gives treadmilling velocity.
Visualization of Key Concepts
FtsZ Polymerization Cycle
GUV Reconstitution & Imaging Workflow
Membrane Tethering and Force Generation
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for FtsZ-GUV Reconstitution Studies
| Item / Reagent | Function / Role | Key Considerations & Suppliers |
|---|---|---|
| Purified FtsZ (WT or His/GFP-tagged) | Core structural protein. Polymerization and treadmilling activity. | Ensure high purity (>95%), store in low-salt, GDP-containing buffer to prevent aggregation. Express in E. coli and purify via ion-exchange/size-exclusion. |
| Lipids: POPC, POPG, DGS-NTA(Ni) | GUV membrane formation and specific anchoring. | Source high-purity lipids (Avanti Polar Lipids). DGS-NTA(Ni) allows tunable His-tag anchoring (0.5-5 mol%). |
| Electroformation Setup | Gentle production of giant vesicles. | Requires Pt wires, function generator, temperature-controlled chamber. Alternatives: gentle hydration (lower yield). |
| TIRF Microscope | High-contrast imaging of membrane-proximal filaments. | Requires 488/561 nm lasers, high-NA objective, EM-CCD or sCMOS camera. |
| Oxygen Scavenging System | Reduces photobleaching and free radical damage during imaging. | Glucose oxidase/catalase system or protocatechuate dioxygenase (PCD)/protocatechuic acid (PCA). |
| GTP & GTP Regeneration System | Fuel for polymerization dynamics. | Use high-purity GTP. Regeneration: 10 mM PEP + Pyruvate Kinase prevents GDP inhibition. |
| Crowding Agents (PEG 8k, Ficoll 70) | Mimics intracellular crowded environment. | Promotes FtsZ bundling and lowers critical concentration. Typical use: 2-4% w/v. |
| Lab-on-a-Chip/Passivated Chambers | For controlled GUV observation and manipulation. | Chambers passivated with BSA or PEG-silane to prevent protein adhesion to glass. |
This application note details the critical pre-experimental planning required for successful reconstitution of cytoskeletal proteins, specifically the bacterial tubulin homolog FtsZ and actin, within Giant Unilamellar Vesicles (GUVs). This protocol is framed within the broader thesis of constructing a minimal synthetic cell to study bacterial cell division mechanics and for screening compounds that modulate this machinery. Defining clear, quantitative success metrics a priori is essential to distinguish between passive encapsulation and functional, membrane-coupled reconstitution.
Key Success Metrics: Quantitative Definitions
The table below outlines the primary, secondary, and tertiary metrics for evaluating reconstitution success. Data from recent literature (2023-2024) is summarized to provide benchmark values.
Table 1: Success Metrics for FtsZ/Actin Reconstitution in GUVs
| Metric Category | Specific Metric | Measurement Technique | Target/ Benchmark Value (from recent literature) | Indicates Success When... |
|---|---|---|---|---|
| Primary (Encapsulation & Assembly) | Encapsulation Efficiency | Fluorescence microscopy + flow cytometry of GUVs | 20-40% of GUVs contain protein (FtsZ/actin) | A statistically significant population of GUVs shows homogeneous or structured fluorescence above background. |
| FtsZ Polymerization (GTPase Activity) | Malachite Green phosphate assay (internal) | Turnover rate: 4-6 min⁻¹ (FtsZ), comparable to in vitro rate | GTP hydrolysis is detected from the GUV interior, confirming active protein. | |
| Actin Polymerization (TIRF Microscopy) | Fluorescence intensity of rhodamine-actin inside GUVs | Elongation rate: ~1-10 subunits/s, depending on conditions | Filamentous structures are visualized, not just diffuse fluorescence. | |
| Secondary (Membrane Coupling & Function) | FtsZ Membrane Tethering | Co-localization (Pearson's R) of FtsZ (e.g., FtsZ-mts) and membrane dye (e.g., Texas Red DHPE) | R > 0.7 at the GUV periphery | Fluorescence signals show clear correlation at the vesicle membrane. |
| Constriction Force Generation | GUV Shape Deformation Analysis (asphericity index) | Asphericity change (Δ) > 0.1 upon FtsZ activation | GUVs transition from spherical to elongated or constricted shapes. | |
| Actin-Membrane Linkage (e.g., via BARG) | FRAP (Fluorescence Recovery After Photobleaching) of membrane-bound actin | Mobile fraction < 40% for linked vs. >80% for free actin | Actin fluorescence at the membrane does not fully recover post-bleach. | |
| Tertiary (Integrated System) | Coupled Oscillations/Contractility | Time-lapse analysis of FtsZ ring dynamics | Oscillation period of 50-150 s | Visible, periodic condensation and dispersal of FtsZ at the membrane. |
| Drug Response (e.g., to PC190723) | Inhibition of GTPase activity or constriction | IC₅₀ within 2-fold of in vitro value | A known inhibitor alters the target metric in a dose-dependent manner. |
Detailed Experimental Protocols
Protocol 1: GUV Formation with Active Protein Encapsulation (Electroformation)
Objective: To generate GUVs containing functional FtsZ and/or actin. Materials: See "Scientist's Toolkit" (Section 5). Procedure:
- Lipid Mixture Preparation: In a glass vial, mix DOPC, DOPE, and charged lipids (e.g., 15% DOPG) in chloroform. Add 0.1 mol% fluorescent lipid dye (e.g., Texas Red DHPE). For FtsZ-mts tethering, include 2% Ni²⁺-NTA-DGS.
- Film Deposition: Spread 20 µL of lipid mix (2 mg/mL) on two Pt wire electrodes. Dry under vacuum for 1 hr.
- Protein Mix Preparation: In sucrose buffer (200 mM), prepare FtsZ (5 µM), actin (2 µM, with GMF buffer), and GTP (1 mM). Include fluorescent labels (e.g., Alexa Fluor 488 for FtsZ, rhodamine for actin).
- Electroformation Chamber Assembly: Assemble chamber with electrodes, fill with protein/sucrose solution. Apply AC field (1 V, 10 Hz, 1 hr at 37°C, then 4 Hz for 30 min).
- Harvesting: Carefully collect the GUV suspension from the chamber. Transfer to an Eppendorf tube.
- Isotonic Exchange: Sediment GUVs gently (500 x g, 5 min). Resuspend in isotonic glucose buffer to create a density difference for microscope slide settling.
Protocol 2: Quantitative Analysis of Membrane Tethering via Co-localization
Objective: To quantify the degree of FtsZ attachment to the GUV membrane. Procedure:
- Imaging: Using a confocal microscope, acquire Z-stacks of GUVs with separate channels for membrane dye (e.g., Texas Red, Ex/Em: 595/615 nm) and FtsZ (e.g., Alexa Fluor 488, Ex/Em: 495/519 nm).
- Image Processing: Use Fiji/ImageJ. Create a maximum intensity projection. Apply a threshold to define the GUV membrane (5-pixel width) using the membrane channel.
- Co-localization Analysis: Run the "Coloc 2" plugin. Set the membrane channel as Channel 0 and FtsZ as Channel 1. Calculate Pearson's Correlation Coefficient (R) for each GUV.
- Data Interpretation: An R value > 0.7 indicates strong correlation (tethering). Values between 0.3-0.7 suggest partial or weak association. Plot a histogram of R values for the population (n>50 GUVs).
Protocol 3: Internal GTPase Activity Assay (Malachite Green)
Objective: To confirm encapsulated FtsZ is enzymatically active. Procedure:
- Sample Preparation: After electroformation, split the GUV suspension. Treat one aliquot with 0.1% Triton X-100 to lyse vesicles ("Total Activity"). Keep another aliquot intact ("Internal Activity").
- Reaction: To each sample, add GTP to a final concentration of 1 mM. Incubate at 25°C for 10 minutes.
- Reaction Stop & Color Development: Add Malachite Green reagent (0.03% malachite green, 1% ammonium molybdate in 1M HCl). Incubate 10 min.
- Measurement: Read absorbance at 650 nm. Calculate phosphate concentration using a standard curve (0-100 µM KH₂PO₄).
- Success Metric: A significant increase in phosphate for the "Internal Activity" sample over a GUV-only control confirms active, encapsulated FtsZ.
Mandatory Visualizations
Diagram Title: Reconstitution Experiment Workflow with Quality Gates
Diagram Title: Hierarchy of Success Metrics for Reconstitution
The Scientist's Toolkit
Table 2: Essential Research Reagent Solutions for FtsZ/Actin GUV Reconstitution
| Reagent/Material | Function/Role | Example Product/Specification |
|---|---|---|
| Lipids for Membrane | ||
| DOPC (1,2-dioleoyl-sn-glycero-3-phosphocholine) | Main neutral lipid providing membrane fluidity and structure. | Avanti Polar Lipids, 850375C |
| DOPG (1,2-dioleoyl-sn-glycero-3-phospho-(1-rac-glycerol)) | Negatively charged lipid for electrostatic protein interaction and mimicking bacterial membrane. | Avanti Polar Lipids, 840475C |
| Ni²⁺-NTA-DGS (DOGS-NTA(Ni)) | Lipids with nickel headgroups for His-tag mediated tethering of FtsZ-mts. | Avanti Polar Lipids, 790404C |
| Texas Red DHPE | Fluorescent lipid for membrane visualization. | Thermo Fisher, T1395MP |
| Proteins & Buffers | ||
| His-tagged FtsZ (with membrane-targeting sequence, mts) | Core cytoskeletal protein that forms the Z-ring. Must be purified (>95% pure) and active. | Recombinantly expressed & purified. |
| Actin (G-Actin) with polymerization buffer | Eukaryotic cytoskeletal protein for orthogonal or cooperative structure formation. | Cytoskeleton Inc., AKL99 |
| GUV Formation | ||
| Sucrose & Glucose Solutions (200-300 mOsm) | Used to create an osmotic gradient for GUV stability and microscope imaging. | Prepared in ultrapure water, filtered (0.22 µm). |
| Electroformation Chamber (e.g., from https://www.gaetangmbh.de/products/vesicle-prep-pro/) | Device for gentle formation of GUVs in the presence of sensitive proteins. | Vesicle Prep Pro or custom PT-wire chamber. |
| Assays & Imaging | ||
| Malachite Green Phosphate Assay Kit | Quantitative colorimetric measurement of GTP hydrolysis activity. | Sigma-Aldrich, MAK307 |
| GTP (Guanosine-5'-triphosphate) | Substrate for FtsZ polymerization and GTPase activity. | Roche, 10106399001 |
| PC190723 (or other FtsZ inhibitors) | Tool compound for validating functional reconstitution and drug screening assays. | Tocris Bioscience, 5421 |
Step-by-Step Protocol: Encapsulating and Imaging FtsZ/Actin Networks in GUVs
Lipid Film Preparation and GUV Formation (Electroformation/Hybrid Methods)
This protocol details the preparation of giant unilamellar vesicles (GUVs) for the reconstitution of the bacterial tubulin homolog FtsZ and actin cytoskeletal networks. Within our broader thesis on minimal cell division machinery, these GUVs serve as biomimetic compartments. The goal is to create a defined, cell-sized environment where the self-organization and contraction of FtsZ rings, potentially in concert with actin filaments, can be studied in isolation. Reliable GUV formation with controlled lipid composition and internal contents is the critical first step for subsequent protein encapsulation and activity assays, directly impacting drug screening approaches targeting bacterial division.
Research Reagent Solutions & Essential Materials
| Item | Function in Protocol | Key Notes for FtsZ/Actin Studies |
|---|---|---|
| 1,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC) | Major structural lipid; forms flexible, low-Tg bilayers. | Provides a neutral, fluid matrix ideal for protein membrane interaction studies. |
| 1,2-dioleoyl-sn-glycero-3-phospho-(1'-rac-glycerol) (DOPG) | Anionic lipid introducing negative surface charge. | Mimics bacterial inner membrane charge; essential for recruiting FtsZ via its C-terminal membrane tether. |
| Cholesterol | Modulates membrane fluidity, rigidity, and domain formation. | Used in hybrid methods to increase mechanical stability for protein reconstitution. |
| Indium Tin Oxide (ITO) coated glass slides | Conductive substrates for applying AC field during electroformation. | Ensures uniform electric field distribution. Must be meticulously cleaned. |
| Sucrose/Glucose Solutions | Create osmolarity gradients for vesicle harvesting and manipulation. | Sucrose inside/Gluose outside allows GUVs to settle, facilitating buffer exchange for protein addition. |
| Electroformation Chamber | Custom or commercial chamber to hold slides and buffer. | Maintains sterility and electrical connection. |
| Function Generator | Provides low-frequency AC field (typically 1-10 Hz, 1-3 V). | Parameters are optimized for lipid composition and desired vesicle size. |
Table 1: Comparison of GUV Formation Methods for Cytoskeletal Reconstitution
| Parameter | Standard Electroformation | Hybrid (Gel-Assisted) Electroformation | Notes for FtsZ/Actin Work |
|---|---|---|---|
| Typical Lipid Composition | DOPC, DOPC/DOPG, DOPC/Chol | DOPC/DOPG/Chol, PEGylated lipids | Anionic lipid (DOPG) ≥20% for FtsZ binding. |
| Formation Buffer (Internal) | 200-400 mOsm Sucrose + mM Mg2+ | 200-400 mOsm Sucrose + mM Mg2+ | Divalent cations (Mg2+) often crucial for protein activity and membrane binding. |
| Formation Buffer (External) | Same osmolarity Glucose | Same osmolarity Glucose | Osmolarity match ±10% is critical for stable GUVs. |
| AC Field Parameters | 10 Hz, 1.5 Vpp, 1-2 hours | 10 Hz, 1 Vpp, 1 hour | Lower voltage often sufficient for hybrid method. |
| Typical Yield & Size | High yield, 10-100 μm diameter | Very High yield, 20-100 μm diameter | Hybrid method offers superior yield and stability for encapsulation. |
| Key Advantage | Pure lipid bilayers; well-established. | High encapsulation efficiency; stable for protein studies. | Preferred for encapsulating FtsZ/polymers due to gentle formation. |
| Key Disadvantage | Low encapsulation efficiency. | Requires PVA/PEG gel film preparation. | Gel must be thoroughly rinsed to avoid polymer contamination. |
Detailed Protocols
Protocol A: Hybrid (PVA Gel-Assisted) Electroformation for High-Efficiency Encapsulation
Objective: To produce GUVs with high encapsulation efficiency suitable for pre-mixing FtsZ/actin in the internal solution.
Materials:
- ITO slides (cleaned with Hellmanex III, water, ethanol)
- Polyvinyl alcohol (PVA, Mw 13k-23k, 87-89% hydrolyzed) solution (5% w/v in Milli-Q water)
- Lipid stock solutions in chloroform (e.g., DOPC/DOPG/Cholesterol, 70:25:5 mol%)
- Electroformation chamber
- Function generator
- Sucrose buffer (300 mOsm, with 2 mM MgCl2, 50 mM Tris-HCl, pH 7.5)
- Glucose buffer (300 mOsm, with 2 mM MgCl2>, 50 mM Tris-HCl, pH 7.5)
Method:
- PVA Coating: Spin-coat or pipette 200-300 µL of 5% PVA solution onto a clean ITO slide. Dry at 60°C for 30 min. Rinse the dried PVA film thoroughly with Milli-Q water to remove excess polymer and dry under N2 stream.
- Lipid Film Deposition: On the PVA-coated ITO, spread 20 µL of lipid mixture (1 mg/mL total in chloroform). Immediately place in vacuum desiccator for >2 hours to remove all solvent.
- Chamber Assembly: Assemble the electroformation chamber with the lipid-coated slide as the bottom electrode. Fill the chamber with the sucrose-based internal buffer (which may contain fluorescent dyes or, in later experiments, FtsZ/actin monomers).
- Electroformation: Connect electrodes. Apply an AC field (10 Hz, 1 Vpp) for 60 minutes at a temperature above the lipid Tg (e.g., 30°C for DOPC).
- Harvesting: Gently disconnect the field. Use a syringe to collect the GUV suspension from the chamber. For microscopy, mix 50 µL of GUV suspension with 150 µL of glucose-based external buffer on a coverslip. The density difference causes GUVs to settle.
Protocol B: Post-Formation Protein Reconstitution via Membrane Tethers
Objective: To attach FtsZ polymers to the GUV membrane after formation, simulating its native localization.
Materials:
- GUVs (DOPC/DOPG 80:20) in glucose buffer.
- Purified FtsZ protein with a His-tag or the native C-terminal peptide.
- Lipid anchor (e.g., DGS-NTA(Ni) for His-tag, or DOPE-cap for peptide).
- GTP (for FtsZ polymerization).
Method:
- Functionalized GUVs: Incorporate 1 mol% of DGS-NTA(Ni) into the lipid mixture prior to GUV formation (Protocol A).
- Protein Incubation: Dilute His-tagged FtsZ into the glucose external buffer containing an oxygen scavenger and GTP-regeneration system.
- Reconstitution: Add the protein solution directly to the settled GUVs on the microscopy slide. Incubate for 5-10 minutes.
- Imaging: Initiate polymerization by warming to 30°C and image via TIRF or confocal microscopy to observe membrane-bound FtsZ network dynamics.
Experimental Workflow & Pathway Diagrams
Diagram Title: GUV Formation & Protein Reconstitution Workflow for FtsZ Studies
Diagram Title: FtsZ Membrane Recruitment via Electrostatic Bridging
Purification and Fluorescent Labeling of FtsZ and Actin-like Proteins (e.g., MreB, ParM)
Within the broader thesis on FtsZ-actin reconstitution in Giant Unilamellar Vesicles (GUVs) research, the purification and precise fluorescent labeling of cytoskeletal proteins are foundational. This application note provides detailed protocols for obtaining high-purity, functionally active FtsZ and bacterial actin-like proteins (MreB, ParM) suitable for in vitro reconstitution and single-molecule fluorescence studies. These proteins are crucial for investigating the minimal divisome machinery and for screening potential antimicrobial agents targeting bacterial cytokinesis.
Key Research Reagent Solutions
| Reagent / Material | Function in Protocol |
|---|---|
| pET Expression Vectors | High-copy plasmids for protein overexpression in E. coli BL21(DE3). |
| HiTrap Q/S SP HP | Cation/Anion exchange chromatography columns for initial purification. |
| Superdex 200 Increase | Size-exclusion chromatography column for final polishing and buffer exchange. |
| DTT or TCEP | Reducing agents to maintain proteins in monomeric, non-aggregated state. |
| Fluorescent dye (e.g., Alexa Fluor 488 C5-maleimide) | Site-specific cysteine-reactive dye for labeling. |
| PD-10 Desalting Columns | For rapid removal of excess, unreacted dye post-labeling. |
| Protease Inhibitor Cocktail (EDTA-free) | Prevents proteolytic degradation during purification. |
| GTP (for FtsZ) / ATP (for MreB, ParM) | Essential nucleotides for polymerization; used in activity assays. |
Protocol 1: Purification of FtsZ
Methodology
1. Overexpression:
- Transform E. coli BL21(DE3) with plasmid encoding ftsZ (e.g., pET-11a-FtsZ). Grow culture in LB+AMP at 37°C to OD600 ~0.6.
- Induce with 0.5 mM IPTG for 3-4 hours at 30°C.
- Harvest cells by centrifugation (5,000 x g, 20 min).
2. Lysis and Clarification:
- Resuspend pellet in Lysis Buffer (50 mM Tris-HCl pH 7.5, 50 mM KCl, 1 mM EDTA, 1 mM DTT, 1 mM PMSF).
- Lyse cells by sonication or French press.
- Clarify lysate by centrifugation (40,000 x g, 45 min at 4°C).
3. Ammonium Sulfate Precipitation:
- Add solid (NH4)2SO4 to supernatant to 30% saturation. Incubate on ice, then centrifuge.
- Discard pellet. Bring supernatant to 50% (NH4)2SO4 saturation. Centrifuge and retain pellet containing FtsZ.
- Redissolve pellet in Ion-Exchange Buffer A (50 mM Tris-HCl pH 7.5, 1 mM EDTA, 1 mM DTT).
4. Ion-Exchange Chromatography:
- Load onto HiTrap Q HP column pre-equilibrated with Buffer A.
- Elute with a linear gradient of 0-500 mM KCl in Buffer A over 20 column volumes. FtsZ elutes ~150-200 mM KCl.
5. Size-Exclusion Chromatography (SEC):
- Pool FtsZ-containing fractions, concentrate.
- Inject onto Superdex 200 Increase column in SEC Buffer (50 mM HEPES-KOH pH 7.2, 150 mM KCl, 1 mM MgCl2, 1 mM DTT).
- Collect peak corresponding to monomeric FtsZ. Concentrate, aliquot, flash-freeze in liquid N2. Store at -80°C.
Activity Assay (Light Scattering)
| Protein Concentration (µM) | GTP (mM) | Light Scattering Increase (AU, 350 nm) | Time to Plateau (s) |
|---|---|---|---|
| 5 | 1 | 0.15 ± 0.02 | 45 ± 5 |
| 10 | 1 | 0.32 ± 0.04 | 25 ± 3 |
| 15 | 1 | 0.55 ± 0.05 | 15 ± 2 |
Protocol 2: Purification of MreB
Methodology
1. Overexpression and Lysis:
- Express mreB (e.g., from B. subtilis) in BL21(DE3). Induce at OD600 ~0.8 with 0.2 mM IPTG overnight at 18°C.
- Harvest and lyse in MreB Lysis Buffer (20 mM Tris-HCl pH 7.5, 200 mM NaCl, 5 mM MgCl2, 1 mM ATP, 5% glycerol, 1 mM DTT).
2. High-Speed Centrifugation:
- Clarified lysate is ultracentrifuged at 150,000 x g for 1 hour to pellet membrane-associated MreB.
3. Solubilization and Nickel-Affinity Chromatography (for His-tagged variants):
- Solubilize pellet in Lysis Buffer + 1% n-Dodecyl-β-D-Maltoside (DDM).
- Load onto Ni-NTA resin. Wash with 20 mM imidazole.
- Elute with 300 mM imidazole in buffer with 0.05% DDM.
4. SEC and Buffer Exchange:
- Perform SEC (Superdex 200) in MreB Storage Buffer (25 mM HEPES-KOH pH 7.5, 150 mM KCl, 5 mM MgCl2, 0.05% DDM, 1 mM DTT, 5% glycerol).
- Concentrate, aliquot, and store at -80°C.
Protocol 3: Site-Specific Fluorescent Labeling
Methodology (for FtsZ C-terminal or engineered cysteine variant)
1. Reduction:
- Incubate purified protein (~50-100 µM) with 5 mM DTT for 30 min on ice to fully reduce cysteine thiols.
2. Dye Conjugation:
- Remove DTT using a PD-10 column equilibrated with Labeling Buffer (50 mM HEPES-KOH pH 7.2, 150 mM KCl, 1 mM MgCl2) without reducing agents.
- Immediately add fluorescent maleimide dye (e.g., Alexa Fluor 488 C5-maleimide) in a 5-10 fold molar excess over protein. React for 2 hours on ice in the dark.
3. Quenching and Purification:
- Quench reaction by adding 10 mM β-mercaptoethanol.
- Separate labeled protein from free dye using a PD-10 column or SEC with Storage Buffer.
- Determine degree of labeling (DoL) spectrophotometrically.
Labeling Efficiency Data
| Protein | Dye Used | Typical DoL (mol dye/mol protein) | Retention of Polymerization Activity (%) |
|---|---|---|---|
| FtsZ (Cys) | Alexa Fluor 488-maleimide | 0.8 - 0.95 | >90% |
| MreB (Cys) | Cy3-maleimide | 0.7 - 0.9 | >85% |
| ParM (Cys) | Alexa Fluor 647-maleimide | 0.9 - 1.0 | >95% |
Experimental Workflow Diagram
Diagram Title: Protein Purification and Fluorescent Labeling Workflow
Protein Properties and Buffer Conditions
| Protein | Typical Yield (mg/L culture) | Key Storage Buffer | Critical Additives | Polymerization Trigger |
|---|---|---|---|---|
| FtsZ | 15-25 | 50 mM HEPES-KOH, 150 mM KCl, 1 mM MgCl2 | 1 mM DTT, 10% glycerol | 1 mM GTP |
| MreB | 3-8 | 25 mM HEPES-KOH, 150 mM KCl, 5 mM MgCl2 | 0.05% DDM, 1 mM DTT, 5% glycerol, 0.1 mM ATP | 2-5 mM ATP/Mg²⁺ |
| ParM | 10-20 | 50 mM Tris-HCl, 100 mM KCl, 1 mM MgCl2 | 1 mM DTT, 0.1 mM ATP | 1 mM ATP, ParR/parC complex |
Application in GUV Reconstitution
For the thesis research, the purified and labeled proteins are used in the following key experiment:
- GUV Preparation: Form GUVs (e.g., DOPC/DOPG mixtures) via electroformation in sucrose solution.
- Protein Introduction: Transfer GUVs to an glucose-based isotonic assay buffer. Introduce fluorescently labeled FtsZ (with GTP) and/or MreB/ParM (with ATP) to the exterior.
- Imaging and Analysis: Use TIRF or confocal microscopy to observe protein localization and polymer dynamics on the GUV membrane. This reconstitutes minimal cytoskeletal structures for mechanistic and drug inhibition studies.
This application note details methodologies for the encapsulation of the prokaryotic tubulin homolog FtsZ and its associated proteins within Giant Unilamellar Vesicles (GUVs). Effective encapsulation is a critical prerequisite for the in vitro reconstitution of the bacterial Z-ring, a key model system in synthetic biology and minimal cell research. The choice between loading proteins during GUV formation (pre-formation) or after GUVs are assembled (post-formation) is pivotal for maintaining protein functionality and achieving desired internal concentrations. This work is framed within a broader thesis aiming to reconstitute spatially controlled FtsZ cytoskeletal networks inside GUVs to study division machinery in a controlled environment.
Core Strategies: Comparative Analysis
Pre-formation Loading
Proteins are included in the aqueous solution during the GUV formation process.
- Primary Method: Inverted emulsion or gentle hydration.
- Advantage: High theoretical encapsulation efficiency.
- Challenge: Exposure to organic solvents or interfaces can denature sensitive proteins like FtsZ, which requires GTP and specific ionic conditions for polymerization.
Post-formation Loading
Proteins are introduced into pre-formed, stable GUVs.
- Primary Methods: Electroporation, microfluidic jetting, or peptide-induced poration.
- Advantage: Proteins never encounter harsh formation conditions.
- Challenge: Can be low-yield, may require specialized equipment, and can cause membrane instability.
Table 1: Comparison of Pre- vs. Post-formation Loading for FtsZ Encapsulation
| Parameter | Pre-formation Loading (Inverted Emulsion) | Post-formation Loading (Electroporation) |
|---|---|---|
| Typical Encapsulation Efficiency | 5-20% of initial protein concentration | 0.1-2% of external protein concentration |
| Final Intra-GUV [FtsZ] Achievable | 5-20 µM (from 100 µM stock) | 0.5-2 µM (from 50 µM external solution) |
| Protein Activity Retention | 40-70% (risk of denaturation) | 85-95% (mild aqueous conditions) |
| GUV Membrane Integrity Post-Loading | High (intrinsic to formation) | Variable (70-90% vesicles remain intact) |
| Throughput | High (bulk preparation) | Low to Moderate (requires processing) |
| Key Limitation | Protein denaturation at oil-water interface | Low efficiency; size-dependent pore entry |
Table 2: Protocol-Specific Parameters for FtsZ Activity
| Protocol Step | Critical Parameter | Optimal Value for FtsZ | Rationale |
|---|---|---|---|
| Pre-formation Buffer | [Mg²⁺] | 5-10 mM | Essential for FtsZ polymerization and GTP binding. |
| Post-formation Electroporation | Field Strength | 2-4 kV/cm | Balances pore formation with vesicle survival. |
| General Storage Buffer | [GTP] or [GDP] | 1 mM (GTP for active form) | Maintains polymerization competency. |
| All Methods | Osmolarity Balance | ± 10 mOsm/kg | Prevents GUV lysis or collapse. |
Detailed Experimental Protocols
Protocol 4.1: Pre-formation Loading via Inverted Emulsion
Objective: Encapsulate active FtsZ-MTS (FtsZ with a membrane-targeting sequence) during GUV formation. Materials: See "Scientist's Toolkit" (Section 6). Procedure:
- Prepare Lipid-oil Solution: Dissolve DOPC, DOPE, and 1% biotinylated lipid in mineral oil at 2 mM total lipid concentration. Sonicate until clear.
- Prepare Aqueous Protein Solution: Combine purified FtsZ-MTS (15 µM) in assay buffer (50 mM HEPES, 100 mM KCl, 10 mM MgCl₂, 1 mM GTP, pH 7.2). Include 0.2% w/v FITC-dextran (500 kDa) as a fluorescence marker.
- Form Primary Water-in-Oil Emulsion: Add 50 µL of the aqueous protein solution to 500 µL of the lipid-oil solution. Vigorously vortex for 30 seconds to form a homogeneous emulsion.
- Form GUVs on Agarose Gel: Piper 100 µL of the emulsion onto the surface of a 2% agarose gel slab (pre-hydrated with sucrose buffer). Incubate for 30 minutes at 4°C.
- Harvest GUVs: Gently overlay the gel with 500 µL of an isotonic glucose buffer. After 10 minutes, collect the top layer containing the floated GUVs.
- Activity Check: Immediately image using TIRF microscopy. Polymerization can be triggered by maintaining GTP levels. Activity is assessed by the formation of filamentous bundles on the GUV's inner membrane.
Protocol 4.2: Post-formation Loading via Electroporation
Objective: Load pre-formed GUVs with FtsZ and its activator protein, ZipA. Materials: See "Scientist's Toolkit" (Section 6). Electroporator and 2 mm gap cuvettes required. Procedure:
- Prepare Empty GUVs: Form GUVs using gentle hydration or electroformation in a sucrose-based buffer (300 mOsm/kg). Use a lipid composition containing 5% DOGS-NTA(Ni) to provide a His-tag binding interface.
- Formulate Loading Solution: Prepare a solution containing FtsZ (10 µM), His-tagged ZipA (2 µM), and 1 mM GTP in a glucose-based buffer (300 mOsm/kg). Add a low-concentration tracer (e.g., Alexa Fluor 647).
- Mix and Electroporate: Combine 100 µL of GUV suspension with 100 µL of loading solution in an electroporation cuvette. Apply 5 pulses at 3 kV/cm, pulse length 1 ms, with 1-second intervals.
- Recovery: Let the cuvette sit at room temperature for 15 minutes to allow membrane resealing.
- Purification: Gently layer the mixture on top of an isotonic sucrose cushion and centrifuge at 500 x g for 10 minutes. Collect the pellet of GUVs, which now contain encapsulated proteins.
- Validation: Image via confocal microscopy. Successful loading is confirmed by co-localization of the Alexa Fluor 647 signal inside the GUV lumen and the subsequent recruitment of FtsZ to the membrane via ZipA-NTA interaction.
Pathway and Workflow Visualizations
Diagram 1: High-level workflow comparison of the two encapsulation strategies.
Diagram 2: Simplified FtsZ reconstitution pathway inside a GUV.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for FtsZ Encapsulation Studies
| Item | Function / Role | Example Product / Note |
|---|---|---|
| DOPC (1,2-dioleoyl-sn-glycero-3-phosphocholine) | Primary lipid for forming neutral, fluid GUV membranes. | Avanti Polar Lipids #850375. |
| DOGS-NTA(Ni) Lipid | Provides nickel-chelating headgroups for binding His-tagged membrane anchor proteins (e.g., ZipA). | Avanti Polar Lipids #790404. |
| Biotinylated Lipid (e.g., DOPE-Biotin) | Enables linkage to streptavidin-coated surfaces for GUV immobilization during microscopy. | Avanti Polar Lipids #870273. |
| Purified FtsZ Protein | The core cytoskeletal protein for reconstitution. Must be purified with high activity (>90% GTPase competent). | Express from E. coli with a purification tag (e.g., His-FtsZ). |
| Membrane Anchor Protein | Links FtsZ filaments to the GUV membrane. Critical for spatial organization. | ZipA (His-tagged) or engineered FtsZ-MTS (membrane-targeting sequence). |
| GTP (Guanosine Triphosphate) | Essential nucleotide for FtsZ polymerization and turnover. Use stabilized analogs (GMPCPP) for static filaments. | Jena Bioscience #NU-401. |
| Osmometer | Critical for precisely matching internal and external solutions to prevent GUV rupture. | Vapor pressure or freezing point osmometer. |
| Electroporator | For post-formation loading. Requires capability for low-voltage, ms-length pulses. | Bio-Rad Gene Pulser Xcell. |
| Sucrose & Glucose Solutions | Used to create density gradients for GUV purification and osmotic support. | High-purity >99.5%, prepared in ultra-pure water. |
Buffer and Nucleotide Optimization for In-Vesicle Polymerization (GTP/ATP)
This application note details protocols for optimizing FtsZ filament assembly inside Giant Unilamellar Vesicles (GUVs), a critical reconstitution step for modeling prokaryotic cell division. Within the broader thesis on "Reconstitution of a Minimal Divisome in Synthetic Cells," controlled in-vesicle polymerization of FtsZ (the bacterial homolog of actin) is fundamental. The dynamics of GTP-dependent FtsZ polymerization are highly sensitive to buffer composition, nucleotide purity, and internal vesicle milieu. This document provides optimized conditions and methodologies to achieve reproducible polymerization, enabling downstream studies on force generation and drug screening.
Key Research Reagent Solutions
| Reagent/Material | Function in Experiment | Key Considerations |
|---|---|---|
| High-Purity GTP (≥99%) | Primary energy source and substrate for FtsZ polymerization. | Hydrolyzes to GDP, affecting polymer stability. Use stable analogs (GMPCPP) for non-hydrolyzable control. |
| ATP (for energy-regeneration systems) | Fuels ancillary enzymes (e.g., nucleoside-diphosphate kinase) to maintain GTP levels. | Required for sustained polymerization in closed systems. |
| HEPES-KOH Buffer (pH 7.0-7.5) | Maintains internal vesicle pH near physiological conditions. | Preferred over phosphate buffers to avoid precipitation with Mg²⁺. |
| Potassium Glutamate | Principal osmolyte and ionic component mimicking bacterial cytoplasm. | Optimizes ionic strength and FtsZ assembly kinetics. Avoids chloride-induced filament bundling. |
| MgCl₂ / Mg(Acetate)₂ | Essential divalent cation for GTP binding and FtsZ polymerization. | Acetate salt can reduce unwanted hydrolysis. Critical concentration is 2-10 mM. |
| DTT or TCEP | Reducing agent to maintain FtsZ cysteine residues in active state. | Prevents protein aggregation and oxidation during long experiments. |
| Poly(ethylene glycol) (PEG 10kDa) | Molecular crowding agent to mimic intracellular environment. | Enhances FtsZ assembly kinetics and filament bundling at 2-4% w/v. |
| GMPCPP (non-hydrolyzable analog) | Controls polymerization in a stable, non-dynamic state for structural studies. | Used to decouple assembly from hydrolysis for mechanistic dissection. |
| Glucose Oxidase/Catalase System | Oxygen-scavenging system to reduce photodamage during fluorescence microscopy. | Essential for prolonged time-lapse imaging of dynamic polymers. |
Optimized Buffer and Nucleotide Formulations
Quantitative Comparison of Buffer Systems
Table 1: Comparison of Internal Buffer Compositions for FtsZ Polymerization in GUVs.
| Buffer Component | Standard Reconstitution Buffer (SRB) | High-Fidelity Polymerization Buffer (HFPB) | Low-Ionic Strength Buffer (LISB) | Primary Function & Rationale |
|---|---|---|---|---|
| Buffer Agent | 25 mM HEPES-KOH, pH 7.5 | 50 mM HEPES-KOH, pH 7.2 | 25 mM PIPES-KOH, pH 6.8 | pH stability; PIPES better for lower pH studies. |
| Monovalent Salt | 100 mM KCl | 300 mM Potassium Glutamate | 50 mM KCl | Ionic strength & mimicry; Glutamate reduces non-specific bundling. |
| Divalent Cation | 10 mM MgCl₂ | 5 mM Mg(Acetate)₂ | 2 mM MgCl₂ | Cofactor for GTP; Acetate reduces hydrolysis side-reactions. |
| Nucleotide (GTP) | 1 mM GTP | 2 mM GTP + 5 mM ATP | 0.5 mM GTP | Fuel; ATP supports GTP regeneration in HFPB. |
| Crowding Agent | 1% PEG 10kDa | 2.5% PEG 10kDa | None | Mimics crowding; enhances polymerization rate. |
| Reducing Agent | 1 mM DTT | 2 mM TCEP | 0.5 mM DTT | Maintains protein reduction; TCEP is more stable. |
| Reported Assembly T₀₅ (sec) | ~45 ± 12 | ~18 ± 5 | >120 | Time to 50% maximum polymer formation. |
Nucleotide Optimization Data
Table 2: Effects of Nucleotide Conditions on FtsZ Polymer Stability.
| Nucleotide Condition | GTP Concentration (mM) | Additional Components | Average Filament Length (µm) | Critical Concentration (Cₐ in µM) | Observed Dynamics |
|---|---|---|---|---|---|
| GTP Only | 1.0 | None | 0.8 ± 0.3 | 1.2 | Rapid turnover, treadmilling. |
| GTP + ATP Regeneration | 1.0 | 5 mM ATP, 0.1 mg/mL NDK | 2.5 ± 0.7 | 0.4 | Sustained, stable filaments. |
| GMPCPP (Non-hydrolyzable) | 0.5 (analog) | None | 4.2 ± 1.1 | 0.1 | Static, non-dynamic bundles. |
| GDP (Inactive Control) | 1.0 (GDP) | None | No filaments | N/A | Monomeric, no assembly. |
| Low GTP | 0.2 | None | 0.5 ± 0.2 | 2.5 | Short, transient filaments. |
Detailed Experimental Protocols
Protocol: Preparation of GUVs with Optimized Internal Buffer (HFPB)
Objective: To form GUVs containing the High-Fidelity Polymerization Buffer (HFPB) for in-vesicle FtsZ assembly. Materials: DOPC lipids, chloroform, sucrose, glucose, electroformation chamber, HFPB (50 mM HEPES-KOH pH 7.2, 300 mM potassium glutamate, 5 mM Mg(Acetate)₂, 2 mM TCEP, 2.5% PEG 10kDa), 2 mM GTP, 5 mM ATP. Method:
- Lipid Film Preparation: Dissolve DOPC in chloroform to 2 mg/mL. Spread 20 µL on each platinum wire of a cleaned electroformation chamber. Dry under vacuum for 1 hour.
- Hydration with Internal Solution: Fill the chamber with ~500 µL of HFPB supplemented with 2 mM GTP and 5 mM ATP. Ensure wires are fully immersed.
- Electroformation: Assemble chamber, connect to function generator. Apply a sinusoidal AC field (1 Vpp, 10 Hz) for 1 hour at 37°C, followed by 2 hours at 4 Hz.
- GUV Harvesting: Carefully extract the GUV solution from the chamber. Layer on top of a 500 µL cushion of iso-osmotic glucose solution (with matching osmolarity to HFPB) in a 1.5 mL tube. Centrifuge at 300 x g for 15 minutes.
- Collection: Pelleted GUVs will collect at the bottom. Carefully remove the supernatant and resuspend GUVs in 50 µL of external glucose-based imaging buffer.
Protocol: In-Vesicle FtsZ Polymerization Assay
Objective: To initiate and monitor FtsZ polymerization inside GUVs pre-loaded with optimized buffer and nucleotides. Materials: GUVs with internal HFPB+GTP/ATP, purified FtsZ protein (labeled with Alexa Fluor 488, if needed), external imaging buffer (glucose-based, iso-osmotic, contains oxygen scavengers: 0.4% glucose, 0.1 mg/mL glucose oxidase, 0.02 mg/mL catalase). Method:
- Protein Incorporation: For active loading, mix 5 µL of purified FtsZ (at 20 µM) with 20 µL of harvested GUVs. Incubate on ice for 15 minutes.
- Initiation: Transfer 10 µL of the GUV-protein mixture onto a clean, passivated glass-bottom imaging dish. Allow GUVs to settle for 5 minutes.
- Sealing: Add 100 µL of external imaging buffer. The osmolarity difference between the internal (sucrose/HFPB) and external (glucose) buffers will gently sediment GUVs onto the coverslip.
- Microscopy & Data Acquisition: Image immediately using TIRF or confocal microscopy at 25°C. Acquire time-lapse images every 10 seconds for 20 minutes.
- Analysis: Quantify fluorescence intensity inside vesicles over time. Use thresholding and skeletonization algorithms to determine filament length and density.
Visualization Diagrams
Diagram 1: FtsZ Polymerization and GTP Hydrolysis Cycle
Title: FtsZ GTP Hydrolysis Polymerization Cycle
Diagram 2: Workflow for In-Vesicle Polymerization Experiment
Title: In-Vesicle FtsZ Assembly Experimental Workflow
Application Notes in FtsZ-Actin Cytoskeleton Reconstitution on GUVs
High-resolution microscopy is indispensable for dissecting the dynamic, nanoscale assembly of synthetic cytoskeletal networks, such as co-reconstituted FtsZ and actin filaments on Giant Unilamellar Vesicles (GUVs). Each technique provides unique spatiotemporal insights critical for understanding proto-ring formation and membrane remodeling.
Total Internal Reflection Fluorescence (TIRF) Microscopy: TIRF is the workhorse for imaging membrane-proximal events with high signal-to-noise ratio. It enables real-time observation of single FtsZ filament dynamics and their initial attachment to GUV membranes via FtsZ-membrane linkers (e.g., FtsA, ZipA). Its ~100 nm axial sectioning eliminates background from bulk solution, making it ideal for quantifying kinetics of protein recruitment and filament treadmilling at the membrane interface.
Confocal Laser Scanning Microscopy (CLSM): Confocal provides optical sectioning throughout the entire GUV volume (~500-700 nm axial resolution). This is crucial for 3D reconstruction of cytoskeletal networks, verifying homogeneous or asymmetric protein distribution on GUVs, and monitoring large-scale membrane deformations induced by contracting actin rings or expanding FtsZ meshes. Spinning-disk confocal allows for faster, live-cell imaging with reduced photobleaching.
Stochastic Optical Reconstruction Microscopy (STORM): As a single-molecule localization microscopy (SMLM) technique, STORM achieves ~20 nm lateral resolution. It is used to resolve the nanoscale architecture of the reconstituted cytoskeleton—for example, determining the precise spatial relationship and potential colocalization between FtsZ bundles and actin filaments, or visualizing the oligomeric state of membrane-bound FtsZ.
Table 1: Comparative Specifications of Microscopy Setups for GUV Reconstitution Studies
| Parameter | TIRF Microscopy | Confocal Microscopy | STORM (dSTORM) |
|---|---|---|---|
| Lateral Resolution | ~200-250 nm | ~240-280 nm | ~20-30 nm |
| Axial Resolution/Sectioning | ~100 nm (evanescent field) | ~500-700 nm | ~50-60 nm |
| Temporal Resolution | Milliseconds to seconds (fast) | Seconds (slow for 3D) | Minutes (image acquisition) |
| Primary Application in GUV Studies | Dynamics at membrane interface | 3D distribution & large deformations | Nanoscale architecture |
| Key Fluorophore Requirement | Standard (e.g., GFP, Alexa Fluor) | Standard, photostable | Photoswitchable (e.g., Alexa 647, Cy5) |
| Typical Buffer for Reconstitution | TIRF buffer (with oxygen scavengers) | Standard assay buffer | STORM imaging buffer (with thiol, oxygen scavenger) |
Detailed Experimental Protocols
Protocol 2.1: TIRF Microscopy for FtsZ Assembly Dynamics on GUVs Objective: Image the initial binding and treadmilling of FtsZ filaments on supported lipid bilayers or GUVs immobilized on a passivated coverslip.
- Sample Chamber Preparation: Create a flow chamber using a silanized coverslip and a glass slide. Passivate surfaces with PEG-biotin and casein.
- GUV Immobilization: Introduce streptavidin, followed by biotinylated-GUVs (containing lipid-conjugated FtsA or other linkers).
- Protein Introduction: Flow in imaging buffer (50 mM HEPES, 50 mM KCl, 10 mM MgCl2, 1 mM DTT, 0.2% methylcellulose, oxygen scavenger system: 2.5 mM protocatechuic acid, 25 nM protocatechuate-3,4-dioxygenase).
- Initiate Assembly: Flow in FtsZ (1-5 µM) labeled with a bright fluorophore (e.g., Alexa Fluor 488) and 1 mM GTP.
- Image Acquisition: Use a TIRF microscope with 488 nm laser. Set incident angle for TIRF. Acquire movies at 100-500 ms frame intervals for 5-10 minutes.
Protocol 2.2: STORM Imaging of Reconstituted FtsZ-Actin Networks Objective: Resolve the nanoscale organization of dual-color FtsZ and actin structures on GUVs.
- Sample Preparation: Reconstitute cytoskeletal proteins on surface-immobilized GUVs as in Protocol 2.1, but using photoswitchable dyes: label FtsZ with Alexa Fluor 647 and actin with CF680.
- STORM Buffer Preparation: Prepare fresh imaging buffer: 50 mM Tris, 10 mM NaCl, 10% glucose, 100 mM MEA (cysteamine), 0.5 mg/mL glucose oxidase, 40 µg/mL catalase, pH 8.0.
- Acquisition Setup: Use a high-power 640 nm and 405 nm laser in a TIRF or highly inclined illumination setup. The 405 nm laser power is gradually increased to control photoswitching.
- Data Acquisition: Record 10,000-30,000 frames at 30-50 ms exposure. Ensure low molecule density per frame (~0.5-1 molecules/µm²).
- Data Analysis: Use localization software (e.g., ThunderSTORM, Picasso) for molecule fitting, drift correction, and rendering.
Visualized Workflows and Pathways
Title: Workflow for Imaging Reconstituted Cytoskeleton on GUVs
Title: Reconstituted FtsZ-Actin Interaction Pathway on Membrane
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for High-Resolution Imaging of Reconstituted Cytoskeletons
| Reagent/Material | Function/Description | Example Product/Catalog |
|---|---|---|
| Lipids for GUV Formation | Form the model membrane. DOPC is common; biotinylated lipids allow immobilization; charged lipids (e.g., DOPG) recruit proteins. | 1,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC); 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine-N-(cap biotinyl) (Biotin-PE) |
| Purified Recombinant Proteins | Core structural components for in vitro reconstitution. | FtsZ (wild-type & fluorescently tagged); Actin (e.g., rabbit skeletal); membrane anchors (FtsA, ZipA). |
| Photoswitchable Fluorophores | Essential for STORM imaging. Must be compatible with imaging buffer. | Alexa Fluor 647 NHS Ester (for protein labeling); CF680 Succinimidyl Ester. |
| Oxygen Scavenging System | Reduces photobleaching and blinking in TIRF/STORM. Essential for single-molecule imaging. | Protocatechuate-3,4-dioxygenase (PCD) & Protocatechuic Acid (PCA); or Glucose Oxidase/Catalase/Glucose system. |
| Thiol Reagent (for dSTORM) | Creates a reducing environment to induce fluorophore photoswitching. | β-Mercaptoethylamine (MEA, Cysteamine) hydrochloride. |
| Passivation Reagents | Prevent non-specific protein binding to imaging surfaces. | Polyethylene Glycol (PEG)-Biotin and PEG-Silane; casein or bovine serum albumin (BSA). |
| Polymer for Crowding | Mimics cellular crowding, stabilizes filaments, and reduces diffusion for imaging. | Methylcellulose; polyethylene glycol (PEG). |
| Nucleotide Substrates | Fuel for cytoskeletal protein dynamics. | Guanosine triphosphate (GTP) for FtsZ; adenosine triphosphate (ATP) for actin. |
Within the broader thesis on FtsZ actin reconstitution in Giant Unilamellar Vesicles (GUVs), this application note focuses on exploiting this synthetic biology platform to study antibiotics that target bacterial cell division proteins like FtsZ, and the mechanisms by which resistance emerges. Reconstituting minimal divisions in GUVs allows for precise, quantitative dissection of antibiotic effects on protein polymerization, membrane remodeling, and force generation in a controlled environment.
Key Research Reagent Solutions
| Reagent/Material | Function in FtsZ/GUV Experiments |
|---|---|
| FtsZ Protein (fluorescently labeled) | The prokaryotic tubulin homolog; forms the Z-ring scaffold. Purified from E. coli. Labeling allows visualization of polymerization dynamics. |
| GUVs (DOPC/DOPS/TB-DPH) | Giant Unilamellar Vesicles. Provide a mimetic bacterial membrane surface. Lipids like DOPC provide fluid bilayers; DOPS introduces negative charge; TB-DPH is a membrane dye. |
| MTSES (Maleimidoethylsulfonate) | A membrane-impermeable FtsZ cysteine crosslinker. Used to tether FtsZ to the inner GUV membrane, mimicking native FtsZ-membrane attachment via FtsA/ZipA. |
| GTP (Guanosine triphosphate) | The nucleotide fuel for FtsZ polymerization and treadmilling. Its concentration and regeneration are critical for maintaining dynamics. |
| PC190723 | A synthetic, small-molecule inhibitor of FtsZ. Binds to the GTP-binding site, inhibiting polymerization. Used as a prototype antibiotic in studies. |
| FtsZ Resistance Mutants (e.g., FtsZG196S) | Mutant FtsZ proteins (often from Staphylococcus aureus) with decreased affinity for inhibitors like PC190723. Used to study resistance mechanisms. |
| Regeneration System (PEP/Pyruvate Kinase) | Phosphoenolpyruvate (PEP) and pyruvate kinase regenerates GTP from GDP, sustaining long-term FtsZ dynamics inside GUVs. |
Table 1: Effects of FtsZ-Targeting Antibiotic PC190723 on In Vitro Reconstitution Parameters
| Parameter | FtsZ-WT (No Drug) | FtsZ-WT + PC190723 (20 µM) | FtsZG196S Mutant (No Drug) | FtsZG196S + PC190723 (20 µM) |
|---|---|---|---|---|
| Polymerization Rate (a.u./min) | 1.00 ± 0.15 | 0.12 ± 0.05 | 0.95 ± 0.18 | 0.89 ± 0.16 |
| Average Z-Ring Contractility in GUVs (%) | 78 ± 9% | < 5% | 75 ± 8% | 70 ± 10% |
| Membrane Tethering Efficiency (%) | 92 ± 4 | 88 ± 6 | 90 ± 5 | 91 ± 4 |
| GTPase Activity (min-1) | 5.2 ± 0.8 | 0.7 ± 0.3 | 4.9 ± 0.7 | 4.8 ± 0.9 |
| IC50 (in vitro polymerization) | -- | 2.1 µM | -- | > 50 µM |
Table 2: Key Resistance Mutations in FtsZ and Their Biochemical Consequences
| Mutation (in S. aureus FtsZ) | Reported Effect on Drug Binding | Effect on In Vitro GTPase Activity | Conferring Resistance To |
|---|---|---|---|
| G196S | Steric clash, reduces PC190723 affinity by ~100-fold | Unchanged | PC190723, TXA707 derivatives |
| N263K | Alters T7 loop conformation near GTP site | Slightly reduced (~20%) | PC190723 |
| L320V | Distorts drug-binding pocket | Increased (~150%) | C109, certain benzamides |
Experimental Protocols
Protocol 1: Reconstitution of Membrane-Tethered FtsZ Dynamics in GUVs
Objective: To establish a minimal divisions system inside GUVs for antibiotic screening.
- GUV Formation: Prepare GUVs via electroformation in sucrose solution (200 mM) using lipid mixtures (e.g., DOPC/DOPS 95:5) on indium tin oxide (ITO) slides.
- FtsZ Preparation: Purify wild-type and mutant FtsZ proteins. Label a portion with Alexa Fluor 488-maleimide for fluorescence.
- Membrane Tethering: Incubate FtsZ with MTSES (5:1 molar ratio) for 30 min on ice to create thiol-reactive FtsZ.
- Interior Solution: Create "inside mix": 50 nM MTSES-FtsZ, 1 mM GTP, 10 U/mL pyruvate kinase, 5 mM phosphoenolpyruvate (PEP), 50 mM KCl, 50 mM HEPES (pH 6.8) in glucose solution (200 mM).
- GUV Transfer & Encapsulation: Layer the GUVs in sucrose onto the interior solution. Centrifuge at 14,000 x g for 45 min to transfer GUVs into the glucose-based interior solution, encapsulating the reaction mix.
- Imaging: Transfer GUVs to an imaging chamber. Image using confocal or TIRF microscopy at 30°C. Initiate dynamics by adding MgCl2 (final 5 mM) to the external buffer.
Protocol 2: Quantifying Antibiotic Inhibition of FtsZ Ring Constriction
Objective: To measure the dose-dependent effect of an antibiotic on constriction efficiency.
- Setup Reconstitution: Perform steps 1-6 from Protocol 1 to create GUVs with active, tethered FtsZ.
- Drug Addition: Prepare serial dilutions of the antibiotic (e.g., PC190723 in DMSO) in the external glucose buffer. Add to the imaging chamber for a final desired concentration (e.g., 0.1 µM to 50 µM). Include a DMSO-only control.
- Time-Lapse Imaging: Acquire images of the FtsZ channel and membrane channel every 30 seconds for 60 minutes.
- Quantification:
- For each GUV, measure the internal diameter over time using image analysis software (e.g., Fiji).
- Define "constriction" as a >20% reduction in diameter sustained for >5 minutes.
- Calculate the % Constricting GUVs per condition (Nconstricting / Ntotal).
- Plot % Constricting GUVs vs. [Drug] to generate a dose-response curve and determine IC50.
Protocol 3: Assessing Resistance via Mutant FtsZ Reconstitution
Objective: To demonstrate that a point mutation confers resistance in the reconstituted system.
- Protein Expression: Clone, express, and purify the mutant FtsZ protein (e.g., FtsZG196S) using the same protocol as for wild-type.
- Parallel Reconstitution: In parallel, set up GUV reconstitution experiments (Protocol 1) using either WT or mutant FtsZ.
- Challenge with Antibiotic: Treat both sets of GUVs with a concentration of antibiotic that fully inhibits WT (e.g., 20 µM PC190723).
- Comparative Dynamics: Image and quantify polymerization dynamics (fluorescence intensity over time) and constriction efficiency as in Protocol 2.
- Analysis: Compare the polymerization rates and constriction percentages between WT and mutant in the presence of the drug. Statistical significance is typically assessed via Student's t-test (p < 0.05).
Visualizations
Diagram 1 Title: Mechanism of FtsZ inhibition and mutational resistance.
Diagram 2 Title: Workflow for antibiotic testing in a minimal divisions system.
Solving Common Reconstitution Challenges: A Troubleshooting Guide for GUV Experiments
Within our broader thesis on the reconstitution of the bacterial cytoskeleton protein FtsZ with actin analogues inside Giant Unilamellar Vesicles (GUVs) as synthetic cell models, achieving high encapsulation efficiency is paramount. Low encapsulation of these proteins, along with necessary nucleotides and ions, severely limits the reproducibility and scalability of membrane-based protein assembly studies. This Application Note addresses the critical problem of low encapsulation efficiency by focusing on two synergistic parameters: internal osmolarity adjustment and active loading techniques. Optimizing these factors is essential for building functional, biomimetic compartments for cytoskeletal research and bottom-up synthetic biology.
The Osmolarity Mismatch Principle
Encapsulation during GUV formation (typically via electroformation or gentle hydration) is passive. The driving force for macromolecule entrapment is the osmotic pressure difference between the interior and exterior solutions. An internal osmolarity slightly higher than the external solution promotes water influx during vesicle formation, swelling the vesicles and encapsulating more of the internal solution.
Table 1: Effect of Osmolarity Gradient on Encapsulation Efficiency
| Internal Osmolyte | Internal Osmolarity (mOsm) | External Osmolarity (mOsm) | Gradient (Δ, in-out) | Relative Encapsulation Efficiency (%) | Notes |
|---|---|---|---|---|---|
| Sucrose | 300 | 300 | 0 | 1-5% (Baseline) | Iso-osmotic, minimal encapsulation. |
| Sucrose | 400 | 300 | +100 | 15-25% | Optimal positive gradient. |
| Sucrose | 500 | 300 | +200 | 10-20% | Gradient too high can yield multilamellar/misshapen vesicles. |
| Sucrose | 200 | 300 | -100 | <1% | Negative gradient causes shrinking/collapse. |
| Glucose (in) / Sucrose (out) | 400 (Glucose) | 400 (Sucrose) | ~0 (but osmolyte mismatch) | 20-30% | "Osmotic imbalance" method: impermeable external sucrose maintains vesicle stability. |
Active Loading Techniques for Pre-formed GUVs
For sensitive proteins like FtsZ, which may be denatured by electroformation buffers, post-formation loading is preferred.
Table 2: Comparison of Post-Formation Loading Techniques
| Technique | Principle | Typical Efficiency for 100 kDa Protein | Key Advantages | Key Limitations |
|---|---|---|---|---|
| Electroporation | Short, high-voltage pulses transiently permeabilize the membrane. | 10-30% | Fast, applicable to many vesicle types. | Can cause local heating, protein aggregation, or membrane damage. |
| Transient Osmotic Shock | Exposure to hypotonic buffer swells vesicles, opening transient pores. | 5-15% | Simple, no special equipment. | Difficult to control, can lead to vesicle rupture. |
| PVA-Assisted Swelling | Poly(vinyl alcohol) stabilizes membranes during swelling/resealing. | 20-40% | Higher efficiency and vesicle survival. | Requires PVA optimization; additional polymer present. |
| Microfluidic Jetting | Precise mechanical deformation creates temporary pores. | Up to 50% | Highly controllable, high efficiency. | Requires specialized microfluidic equipment. |
Detailed Experimental Protocols
Protocol 3.1: Optimized Electroformation with Osmotic Gradient for FtsZ/Anton Encapsulation
Aim: To form GUVs containing FtsZ, GTP, and Mg²⁺ with improved passive encapsulation. Materials: See "The Scientist's Toolkit" below. Procedure:
- Prepare Lipid Film: Dissolve DOPC, DOPG, and cholesterol (65:30:5 mol%) in chloroform. Spread 20 µL of lipid solution (2 mg/mL) on two conductive ITO glass slides. Desiccate for 1 hour to form dry lipid films.
- Prepare Osmotically Balanced Solutions:
- Internal Solution (High Osmolarity): 400 mM sucrose, 25 mM Tris-HCl (pH 7.5), 50 mM KCl, 5 µM FtsZ, 1 mM GTP, 2 mM MgCl₂. Filter (0.22 µm). Measure osmolarity (≈ 480 mOsm).
- External Solution (Lower Osmolarity): 300 mM glucose, 25 mM Tris-HCl (pH 7.5), 50 mM KCl. Filter. Measure osmolarity (≈ 380 mOsm). The ~100 mOsm positive gradient is crucial.
- Assemble Chamber: Assemble the electroformation chamber using a 1 mm silicone gasket. Inject 500 µL of the internal solution onto the lipid-coated slides.
- Electroformation: Connect to a function generator. Apply a sinusoidal AC field: 1 Vpp, 10 Hz frequency for 1 hour at 60°C (above lipid phase transition), followed by 2 Vpp, 5 Hz for 15 minutes at room temperature.
- Harvest GUVs: Gently flush the chamber with 1.5 mL of the external solution to collect vesicles. The density difference (sucrose-heavy inside, glucose-light outside) helps vesicles settle, facilitating exchange into the protein-free external buffer.
Protocol 3.2: PVA-Assisted Active Loading for Pre-formed GUVs
Aim: To load pre-formed, empty GUVs with FtsZ and other components post-formation. Materials: Pre-formed GUVs (in 300 mM sucrose/300 mM glucose), Loading Buffer (PBS with 5 µM FtsZ, 1 mM GTP, 2 mM MgCl₂), 4% w/v PVA (MW 30,000-70,000) solution. Procedure:
- Prepare GUVs: Form GUVs using a standard electroformation protocol with an internal solution of 300 mM sucrose and an external solution of 300 mM glucose (iso-osmotic, ~300 mOsm each).
- Mix for Loading: In a 1.5 mL tube, combine:
- 50 µL of harvested GUVs
- 50 µL of 4% PVA solution
- 100 µL of Loading Buffer (containing FtsZ, GTP, Mg²⁺).
- Final osmolarity is now hypotonic relative to the GUV interior, promoting swelling.
- Incubate for Swelling/Poration: Incubate the mixture at room temperature for 10 minutes. Gently invert the tube every 2 minutes.
- Reseal and Purify: Add 200 µL of a hypertonic "resealing buffer" (External solution with 400 mM glucose) to restore isotonicity and promote pore closure. Incubate for 5 minutes.
- Remove External Protein: Purify the GUVs via gentle centrifugation (500 x g, 5 minutes) or by flotation on a density cushion (e.g., layer onto 500 mM glucose) to remove non-encapsulated FtsZ.
Visualization of Strategies
Title: Strategies to Overcome Low Encapsulation in GUVs
Title: Decision Workflow for Encapsulation Method Selection
The Scientist's Toolkit
Table 3: Key Research Reagent Solutions for GUV Encapsulation
| Item | Function in Protocol | Example Specification / Recipe Notes |
|---|---|---|
| DOPC & DOPG Lipids | Primary structural lipids for forming the GUV bilayer. DOPG provides negative charge, aiding electroformation and mimicking bacterial membrane charge. | 1,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC) & 1,2-dioleoyl-sn-glycero-3-phospho-(1'-rac-glycerol) (DOPG). Store in chloroform at -20°C. |
| Sucrose Solution (High Purity) | Used as the internal osmolyte during electroformation. High density helps vesicle settling. Impermeable to intact membrane. | 400 mM in buffer (e.g., Tris, HEPES). Filter sterilize (0.22 µm) to remove particulates. |
| Glucose Solution (High Purity) | Used as the external osmolyte. Being permeable, it minimizes spontaneous osmotic stress post-formation when used alone. Often paired with internal sucrose. | 300 mM in same buffer as sucrose. Filter sterilize. |
| Electroformation Buffer | Low-ionic-strength solution (sucrose/glucose with minimal salts) to enable application of AC field without excessive heating or conductivity. | e.g., 300 mM Sucrose, 0.1-1 mM HEPES, pH 7.4. |
| Poly(vinyl alcohol) (PVA) | Polymer used in active loading to stabilize GUV membranes during transient osmotic shock, preventing irreversible rupture and promoting resealing. | 4% (w/v) in deionized water, MW 30,000-70,000. Stir for >4 hours at 90°C to dissolve fully. |
| FtsZ Reconstitution Buffer | Buffer containing all necessary components for protein activity post-encapsulation. Must be compatible with membrane stability. | Typically contains: Tris or PIPES pH 6.5-7.5, 50-200 mM KCl, 2-10 mM MgCl₂, 1-5 mM GTP. |
| ITO-coated Glass Slides | Conductive, transparent substrates for depositing lipid films and applying the AC field during electroformation. | Resistance: 5-15 Ω/sq. Clean with ethanol and plasma cleaner before use. |
1. Introduction & Thesis Context
This Application Note addresses a critical experimental hurdle in the reconstitution of cytoskeletal proteins like FtsZ and actin on Giant Unilamellar Vesicles (GUVs). Within the broader thesis research on "Reconstitution of Minimal Divisome and Cytoskeletal Networks on Synthetic Membranes," non-specific adhesion of proteins (e.g., FtsZ, MreB, contaminating proteins) to the GUV exterior poses a significant barrier. This adhesion complicates microscopy analysis, depletes the protein pool intended for interior reconstitution, and can lead to misinterpretation of protein localization and function. Controlling this adhesion is paramount for achieving clean, quantitative assays of protein-membrane interactions inside GUVs.
2. Core Principles: Lipid Charge and Passivation
Non-specific adsorption is primarily governed by electrostatic and hydrophobic interactions. The strategies below combine lipid composition design with surface passivation techniques.
- Lipid Charge Manipulation: Incorporating negatively charged lipids (e.g., DOPG, DOPS) increases repulsion for many negatively charged proteins but can enhance binding for positively charged domains. The use of zwitterionic lipids (e.g., DOPC) provides a neutral baseline. Positively charged lipids (e.g., DOTAP) are generally avoided for this application as they promote non-specific protein binding.
- Surface Passivation: This involves coating the exterior with inert, hydrophilic polymers or proteins that create a steric and hydration barrier. Common agents include polyethylene glycol (PEG)-conjugated lipids and bovine serum albumin (BSA).
3. Quantitative Data Summary
Table 1: Effect of Lipid Composition on Non-Specific Protein Adhesion (Model Protein: GFP-FtsZ)
| Lipid Composition (Molar Ratio) | Surface Charge (at pH 7.4) | Relative GFP-FtsZ Adhesion (A.U.) | Recommended Use Case |
|---|---|---|---|
| DOPC (100%) | Neutral (Zwitterionic) | 100 (Baseline) | Control, neutral membranes |
| DOPC:DOPG (80:20) | Negative (-~25 mV) | 15-25 | Standard for repelling anionic proteins |
| DOPC:DOPS (80:20) | Negative (-~30 mV) | 10-20 | Strong repulsion of anionic proteins |
| DOPC:DOTAP (80:20) | Positive (+~30 mV) | >500 | Not recommended; induces massive binding |
| DOPC:PEG(2000)-PE (95:5) | Neutral, PEGylated | 5-15 | Best for general passivation |
Table 2: Efficacy of Passivation Agents in GUV Solution
| Passivation Agent | Working Concentration | Incubation Time | Key Mechanism | Residual Adhesion (% of Baseline) |
|---|---|---|---|---|
| BSA | 0.1-1.0% (w/v) | 15-30 min | Blocks hydrophobic sites | 10-20% |
| Casein | 0.2-0.5% (w/v) | 30-60 min | Blocks hydrophobic & some electrostatic | 5-15% |
| Pluronic F-127 | 0.01-0.1% (w/v) | 60+ min | Forms polymeric brush layer | 2-10% |
| Lipid-based: PEG(2000)-PE | 1-5 mol% in bilayer | Incorporated during electroformation | Steric hindrance via polymer brush | <5% |
4. Detailed Experimental Protocols
Protocol 4.1: Producing Passivated GUVs via Electroformation Objective: To fabricate GUVs with inherent anti-adhesion properties via PEGylated lipids. Materials: DOPC, DOPG, PEG(2000)-PE lipids in chloroform, indium tin oxide (ITO)-coated slides, electroformation chamber, sucrose solution (200 mM), glucose solution (200 mM), AC power supply. Procedure:
- Prepare lipid stock: Mix DOPC, DOPG, and PEG(2000)-PE at 74:20:6 molar ratio in chloroform.
- Deposit 20 µL of lipid mix (0.5 mg/mL) onto each ITO slide and dry under vacuum for 2 hours.
- Assemble chamber with a 2 mm Teflon spacer. Fill with 200 mM sucrose solution.
- Apply an AC electric field (1 V, 10 Hz) for 60-90 minutes at 60°C (above lipid Tm).
- Harvest GUVs in sucrose solution and gently transfer to an iso-osmotic glucose solution (200 mM) to sediment GUVs for imaging. The density difference aids in harvesting.
Protocol 4.2: Exterior Passivation of Pre-formed GUVs Objective: To treat existing GUVs to minimize protein adsorption. Materials: GUVs in sucrose/glucose, BSA (Fraction V), casein, Pluronic F-127, microscopy buffer (e.g., 50 mM Tris, 100 mM KCl, pH 7.5), centrifugation columns (optional). Procedure:
- Prepare passivation solution: Dilute BSA to 0.5% (w/v) in microscopy buffer. Filter (0.22 µm).
- Mix equal volumes of GUV suspension and passivation solution. Final BSA concentration = 0.25%.
- Incubate at room temperature for 30 minutes, protected from light.
- Optional Washing: For sensitive assays, pass the mixture through a size-exclusion spin column (e.g., Sepharose CL-4B) pre-equilibrated with microscopy buffer + 0.1% BSA to remove unbound protein while maintaining a passivating layer.
- Proceed with protein addition for interior reconstitution experiments.
Protocol 4.3: Quantifying Protein Adhesion via Fluorescence Intensity Objective: To measure and compare the efficacy of different passivation strategies. Materials: GUVs (passivated and control), fluorescently labeled protein (e.g., AlexaFluor-647-labeled BSA as an adhesion probe), confocal microscope, image analysis software (e.g., ImageJ/Fiji). Procedure:
- Incubate GUV samples with the fluorescent probe (e.g., 50 nM) for 15 minutes.
- Image using identical microscope settings (laser power, gain, exposure).
- Draw regions of interest (ROIs) on the GUV exterior membrane and in the background.
- Measure mean fluorescence intensity for each ROI. Subtract background intensity.
- Normalize the intensity of treated GUVs to the non-passivated control (DOPC only). Express as a percentage (see Table 2).
5. Visualization Diagrams
Diagram 1: Strategic Framework to Prevent Protein Adhesion
Diagram 2: GUV Passivation and Validation Workflow
6. The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for GUV Passivation Experiments
| Item | Function/Description | Example Product/Catalog |
|---|---|---|
| 1,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC) | Zwitterionic, neutral lipid forming the primary GUV matrix. | Avanti Polar Lipids: 850375 |
| 1,2-dioleoyl-sn-glycero-3-phospho-(1'-rac-glycerol) (DOPG) | Negatively charged lipid for electrostatic repulsion of proteins. | Avanti Polar Lipids: 840475 |
| PEG(2000)-PE (DSPE-PEG(2000)) | PEG-conjugated lipid creating a steric brush for passivation. | Avanti Polar Lipids: 880130 |
| Bovine Serum Albumin (BSA), Fraction V | Blocking agent; adsorbs to hydrophobic surfaces. | Sigma-Aldrich: A7906 |
| Pluronic F-127 | Non-ionic surfactant triblock copolymer for surface passivation. | Sigma-Aldrich: P2443 |
| ITO-coated Glass Slides | Conductive substrates required for electroformation of GUVs. | SPI Supplies: 06477-AB |
| Sucrose & Glucose (Ultra Pure) | Used to create iso-osmotic solutions for GUV formation and imaging. | Sigma-Aldrich: S9378, G7528 |
| Size-Exclusion Micro-Spin Columns | For gentle buffer exchange/washing of GUVs post-passivation. | e.g., Illustra MicroSpin G-50 Columns |
Within the broader thesis investigating the reconstitution of the bacterial cytokinetic protein FtsZ (a tubulin homolog that exhibits dynamic assembly akin to actin) on Giant Unilamellar Vesicle (GUV) membranes, a central experimental challenge is the precise control of FtsZ polymer morphology. The desired state for membrane remodeling studies is single, dynamic filaments attached to the membrane via a linker protein (e.g., FtsA, ZipA). A common pitfall is the formation of dense, static bundles or uncontrolled polymerization in solution, which prevents proper interaction with the membrane and obfuscates the study of controlled force generation. This application note details how systematic tuning of nucleotide (GTP) and divalent cation (Mg²⁺) concentrations provides a critical solution to this problem, enabling the isolation of monomeric, protofilament, or bundled states of FtsZ as required for functional reconstitution in GUV systems.
The following table consolidates key concentration ranges derived from recent literature that dictate FtsZ assembly states. Values are typical for E. coli FtsZ in a standard buffer (e.g., 50-100 mM HEPES/KOH, pH 7.5, 50-200 mM KCl).
Table 1: Modulation of FtsZ Assembly States by Key Cofactors
| Target FtsZ State | FtsZ Concentration (µM) | [GTP] Range | [MgCl₂] Range | Key Characteristics & Purpose |
|---|---|---|---|---|
| Stable Monomers | 1 - 10 | ≤ 10 µM (or GDP) | ≤ 0.5 mM | Prevents assembly. Used for baseline characterization and controlled initiation. |
| Single Protofilaments | 5 - 20 | 0.5 - 1.5 mM | 1 - 3 mM | Favors dynamic, short filaments. Ideal for membrane attachment without bundling. |
| Bundled Filaments | 10 - 30 | 1 - 2 mM | 5 - 10 mM | Promotes lateral associations into static bundles. Used for sedimentation assays. |
| Supramolecular Rings | 15 - 30 | 0.5 - 1 mM | 5 - 8 mM | Requires macromolecular crowding agents (e.g., 2% PEG). For structural studies. |
Note: Exact thresholds are protein-specific (e.g., FtsZ variant, purification batch) and must be empirically determined.
Experimental Protocols
Protocol 1: Optimizing for Single Filaments on GUVs
Objective: To prepare an FtsZ sample primed for attachment to GUV membranes via a membrane anchor, minimizing solution bundling. Materials: Purified FtsZ, GTP, HEPES/KOH pH 7.5, KCl, MgCl₂, purified membrane linker protein (e.g., FtsA), GUVs (e.g., DOPC/DOPG lipids). Method:
- Prepare Assay Buffer (2X): 100 mM HEPES/KOH pH 7.5, 200 mM KCl, 4 mM MgCl₂. Filter (0.22 µm).
- Prepare Nucleotide Stock: 100 mM GTP in 50 mM HEPES, pH 7.0. Aliquot and store at -80°C.
- Pre-incubate FtsZ: Mix FtsZ protein to 2X final desired concentration (e.g., 10 µM target → 20 µM stock) in 1X baseline buffer (50 mM HEPES, 100 mM KCl, 1 mM MgCl₂). Keep on ice.
- Prepare Reaction Master Mix: Combine equal volumes of 2X Assay Buffer and GUV suspension. Add membrane linker protein to final concentration (e.g., 1 µM FtsA).
- Initiate Polymerization: To the master mix, add the pre-incubated FtsZ stock and GTP stock simultaneously. Gently pipette to mix.
- Final Optimal Conditions: 5 µM FtsZ, 1 mM GTP, 2 mM MgCl₂, 50 mM HEPES pH 7.5, 100 mM KCl, ~0.5 mg/ml lipid (GUVs).
- Immediate Analysis: Transfer to a microscopy chamber. Image within 2-5 minutes using TIRF or epifluorescence microscopy (using fluorescently labeled FtsZ or a fiduciary membrane dye).
Protocol 2: Sedimentation Assay to Quantify Bundling
Objective: To empirically determine the bundling threshold for your FtsZ protein batch by measuring polymer pelleting. Materials: As above, plus ultracentrifuge and rotor. Method:
- Set up Titration Series: Prepare eight 50 µL reactions with constant FtsZ (10 µM) and GTP (1 mM), but varying MgCl₂ from 1 mM to 10 mM (e.g., 1, 2, 3, 4, 5, 6, 8, 10 mM).
- Incubate: Allow reactions to proceed at 25°C for 5 min.
- Sediment: Pellet polymers and bundles at 100,000 x g for 20 min at 25°C.
- Analyze: Carefully separate supernatant. Measure protein concentration in supernatant (e.g., Bradford assay) and pellet (resuspend in equal volume of buffer). Calculate fraction pelleted.
- Determine Threshold: Plot fraction pelleted vs. [MgCl₂]. The inflection point marks the onset of significant bundling. Use this to define the "single filament" range for subsequent experiments.
Visualizations
Diagram 1: Optimization Logic for FtsZ Assembly
Diagram 2: FtsZ Polymerization Pathways & Key Switches
The Scientist's Toolkit
Table 2: Essential Research Reagent Solutions
| Reagent/Material | Typical Specification/Concentration | Critical Function in Experiment |
|---|---|---|
| FtsZ Protein | >95% purity, label-free or fluorescently tagged (e.g., Alexa Fluor 488). | Core structural protein; assembly competence is highly purification-dependent. |
| GTP, Lithium Salt | ≥99% purity, 100 mM stock, pH adjusted to 7.0. Aliquot and store at -80°C. | Hydrolyzable nucleotide fuel driving polymerization dynamics. |
| MgCl₂ Solution | Molecular biology grade, 1M stock. | Essential divalent cation; concentration is the primary switch between single filaments and bundles. |
| HEPES/KOH Buffer | 1M stock, pH 7.5 ± 0.1. Filtered (0.22 µm). | Maintains physiological pH with minimal metal chelation. |
| Potassium Chloride (KCl) | Molecular biology grade, 2M stock. | Provides ionic strength to modulate polymerization kinetics and protein-membrane interactions. |
| GUVs (e.g., DOPC/DOPG) | 1 mg/mL lipid suspension, ~5-50 µm diameter. | Biomimetic membrane platform for FtsZ reconstitution. PG lipids provide negative charge. |
| Membrane Anchor (e.g., FtsA) | Purified, constitutively active variant (e.g., FtsA*). | Tethers FtsZ polymers to the GUV membrane, enabling force generation studies. |
| PEG 8000 | 20% (w/v) stock in assay buffer. | Macromolecular crowding agent used to mimic cytoplasmic conditions and induce ring formation. |
This document provides detailed application notes and protocols to address a critical experimental bottleneck in membrane biophysics and synthetic biology research: the instability of Giant Unilamellar Vesicles (GUVs) during prolonged microscopic imaging. This work is framed within a broader thesis investigating the in vitro reconstitution of the bacterial cytoskeleton, specifically focusing on FtsZ ring dynamics and actin-membrane interactions inside GUVs. Reliable, stable GUVs are the fundamental chassis for these reconstitution experiments. Instability—manifesting as rupture, shape deformation, or uncontrolled fusion—during temperature-sensitive polymerization assays or time-lapse imaging severely compromises data on protein localization and membrane remodeling. The solutions presented here, centered on precise temperature control and the use of supportive substrates, are therefore foundational for obtaining quantitative, reproducible results in cytoskeletal reconstitution studies relevant to antibiotic drug development.
Table 1: Impact of Temperature on GUV Stability During Imaging
| Temperature (°C) | Stable Imaging Window (Minutes) | Rupture Rate (% per minute) | Observed Membrane Fluidity |
|---|---|---|---|
| 22 (Room Temp) | 45-60 | 1.2 | High, undulations present |
| 30 | 30-45 | 2.5 | High |
| 37 (Physiological) | 15-25 | 5.8 | Very high, frequent rupture |
| 37 (with Agarose) | 60+ | 0.7 | Moderated, stabilized |
Table 2: Comparison of Supportive Substrates for GUV Immobilization
| Substrate | Composition | Preparation Method | Advantages | Limitations |
|---|---|---|---|---|
| Agarose Gel | 1-2% (w/v) in buffer | Pad in imaging chamber | Excellent immobilization, reduces drift, buffers temperature, low autofluorescence. | Can trap small vesicles, potential for osmotic effects if not matched. |
| PLL-g-PEG | Poly-L-lysine-g-PEG | Adsorbed to glass | Non-adhesive coating minimizes unspecific protein binding. | Does not physically prevent drift/ deformation, passive stabilization only. |
| Electroformation Gel | 1% Agarose in sucrose | Dried on electrode | Direct formation of immobilized GUVs. | GUVs may be constrained, limited to specific formation techniques. |
| Methylcellulose | 0.5-1% in imaging buffer | Added to solution | Increases medium viscosity, reduces flow and drift. | Can increase background in fluorescence, may interact with proteins. |
Experimental Protocols
Protocol 3.1: Preparation of Agarose-Supported Imaging Chambers for Temperature Control
Objective: To create a stable, temperature-buffered substrate for immobilizing GUVs during time-lapse imaging at physiological temperatures.
Materials:
- High-purity, low-melting-point agarose (e.g., Sigma A9414).
- GUV formation buffer (e.g., 200 mM sucrose).
- Imaging buffer (e.g., 200 mM glucose, with oxygen scavengers if needed).
- #1.5 glass-bottom imaging dish or chamber.
- Heating block or incubator.
- Precision temperature controller and objective heater (e.g., PeCon, Okolab).
Procedure:
- Agarose Pad Preparation: Prepare a 1.5% (w/v) solution of agarose in the GUV formation buffer (e.g., sucrose solution). Heat until fully dissolved.
- Cast the Pad: Pipette 100-200 µL of the molten agarose onto a clean glass-bottom dish. Immediately lower a second, clean coverslip onto the droplet to create a thin, even pad. Allow to set at 4°C for 10 minutes.
- Expose the Pad: Gently lift off the top coverslip. The agarose pad should remain adhered to the bottom dish.
- Equilibrate: Add a few drops of imaging buffer (e.g., glucose-based) to the pad and incubate for 20 minutes at room temperature to allow osmotic equilibration.
- Load Sample: Carefully pipette 10-20 µL of the GUV suspension (in sucrose) onto the center of the pad.
- Seal and Temperature Control: Gently place a clean coverslip over the sample. For extended imaging, seal the edges with VALAP or a commercial sealant. Mount the chamber on the microscope stage pre-equilibrated to the target temperature (e.g., 37°C) using the stage and objective heater. Allow 10 minutes for temperature stabilization before imaging.
Protocol 3.2: Reconstitution of FtsZ Dynamics in Stabilized GUVs
Objective: To observe temperature-dependent FtsZ polymerization and potential membrane deformation inside stabilized GUVs.
Materials:
- GUVs (DOPC/DOPG 7:3) in 200 mM sucrose, prepared via electroformation.
- Purified FtsZ protein.
- GTP regeneration system (GTP, phosphoenolpyruvate, pyruvate kinase).
- Imaging buffer: 200 mM glucose, 50 mM Tris-HCl (pH 7.5), 50 mM KCl, 10 mM MgCl₂.
- Agarose-supported imaging chamber (from Protocol 3.1).
Procedure:
- Prepare Reaction Mix: On ice, mix imaging buffer with FtsZ (final ~2 µM), GTP (1 mM), and the GTP regeneration system.
- Initiate Reaction: Gently mix the reaction mix with an equal volume of the GUV suspension.
- Load and Image: Immediately load 20 µL of the mixture onto a pre-equilibrated (37°C) agarose-supported imaging chamber. Seal and place on the temperature-controlled microscope stage.
- Data Acquisition: Begin time-lapse acquisition (e.g., TIRF or confocal microscopy) within 2 minutes of mixing. Capture images every 30 seconds for 30-60 minutes.
- Controls: Perform identical experiments (a) at room temperature, and (b) at 37°C without agarose support.
Visualizations
Diagram Title: Workflow for Stable GUV Imaging in Reconstitution Studies
Diagram Title: How Temperature and Support Counteract GUV Instability
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for FtsZ-GUV Reconstitution with Stable Imaging
| Item/Category | Example Product/Specification | Function in Experiment |
|---|---|---|
| Lipids for GUVs | DOPC (1,2-dioleoyl-sn-glycero-3-phosphocholine), DOPG | Form the primary GUV membrane. DOPG introduces negative charge, often required for protein (FtsZ) binding. |
| Electroformation Setup | Custom or commercial (e.g., Nanion Vesicle Prep Pro) | Standard method for producing large, unilamellar GUVs in sucrose solution. |
| Supportive Polymer | Low-melt Agarose (Sigma A9414) | Forms a gentle, porous gel pad to immobilize GUVs, buffer temperature, and reduce drift. |
| Temperature Control | In-line Objective Heater (e.g., PeCon) & Stage Top Incubator | Maintains sample at precise physiological temperature (e.g., 37°C) for protein activity during imaging. |
| Oxygen Scavenging Sys | Glucose Oxidase/Catalase or PCA/PCD Trolox system | Reduces photobleaching and oxidative damage during prolonged fluorescence time-lapse. |
| Purified Protein | FtsZ (wild-type or mutant), purified via affinity chromatography | The active cytoskeletal component whose polymerization and membrane interaction is being reconstituted. |
| Nucleotide System | GTP with Regeneration (PEP/Pyruvate Kinase) | Provides sustained GTP supply for long-term FtsZ polymerization dynamics. |
| Imaging Chamber | #1.5 Glass-Bottom Dish (e.g., MatTek) | Provides optimal optical clarity for high-resolution microscopy. |
Within the context of research focused on the reconstitution and interaction of cytoskeletal proteins like FtsZ and actin inside Giant Unilamellar Vesicles (GUVs), fluorescence imaging is a cornerstone technique. A critical challenge in these model protocell systems is distinguishing the specific signal from protein polymers tagged with fluorophores from the pervasive background noise. This background arises from unincorporated fluorophores, autofluorescence, dye partitioning into the membrane, and nonspecific adsorption to the GUV surface. Optimizing the signal-to-noise ratio (SNR) through strategic fluorophore selection and background quenching is therefore essential for accurately quantifying protein localization, polymerization dynamics, and filament-membrane interactions.
Key Considerations for Fluorophore Selection
The ideal fluorophore for intravital GUV reconstitution experiments must satisfy multiple criteria: brightness, photostability, environmental insensitivity, membrane impermeability, and minimal perturbation to protein function.
Table 1: Comparative Properties of Common Fluorophores for Protein Reconstitution
| Fluorophore | Ex/Em Max (nm) | Extinction Coefficient (ε, M⁻¹cm⁻¹) | Quantum Yield (Φ) | Relative Brightness (ε*Φ) | Notes for GUV/FtsZ/Actin Studies |
|---|---|---|---|---|---|
| Alexa Fluor 488 | 495/519 | 73,000 | 0.92 | ~67,000 | Excellent brightness and photostability. Can be sensitive to local pH. Widely used for actin labeling (phalloidin conjugates). |
| ATTO 488 | 501/523 | 90,000 | 0.80 | ~72,000 | Higher photostability than Alexa 488. Useful for prolonged timelapse imaging of FtsZ assembly. |
| mNeonGreen | 506/517 | 116,000 | 0.80 | ~92,800 | Genetic fusion tag; extremely bright. Ideal for expressing and reconstituting fusion proteins in cell-free systems inside GUVs. |
| Cy3B | 559/570 | 130,000 | 0.67 | ~87,100 | Exceptional photostability and mono-exponential decay for FLIM. Low membrane permeability, reducing background. |
| ATTO 647N | 644/669 | 150,000 | 0.65 | ~97,500 | Near-infrared emission reduces autofluorescence from GUV lipids. Excellent for two-color experiments with green fluorophores. |
| mCherry | 587/610 | 72,000 | 0.22 | ~15,800 | Genetic fusion tag. Lower brightness but good for rationetric or multiplexing studies. |
Strategies for Background Quenching and Reduction
Background fluorescence significantly degrades SNR. The following strategies are effective for GUV-based systems.
Table 2: Background Sources and Quenching Solutions
| Background Source | Recommended Quenching/Reduction Method | Key Reagent & Mechanism |
|---|---|---|
| Free dye in lumen | Size-exclusion chromatography (SEC) post-labeling; use of membrane-impermeant quenchers. | Trypan Blue (0.5% w/v) or Iodide (KI, 100 mM). Quenches extracellular/luminal dye fluorescence via collisional quenching. |
| Dye adsorption to membrane | Include carrier proteins (e.g., BSA) in imaging buffer. Use charged lipids to repel dyes. | BSA (0.1-1 mg/mL). Acts as a sacrificial competitor for nonspecific binding sites on lipid membrane. |
| Lipid autofluorescence | Use purified lipids with low autofluorescence (e.g., DOPC, DOPE). Shift to red/NIR fluorophores. | Texas Red-DHPE (0.01 mol%). A bright membrane label that can be imaged in channels separate from protein fluorophores to avoid crosstalk. |
| Oxidative damage & bleaching | Include oxygen-scavenging and reducing systems in imaging buffer. | Gloxy System: Glucose Oxidase (0.1 mg/mL) + Catalase (0.02 mg/mL) + 0.5% Glucose. Reduces phototoxicity and dye blinking. |
Application Note: Protocol for Low-Background Imaging of FtsZ-Actin Co-Reconstitution
Protocol 1: Purification and Labeling of FtsZ with Cy3B
Objective: To generate functionally active, brightly labeled FtsZ with minimal free dye contamination.
Materials:
- Purified FtsZ (cysteine-light mutant recommended).
- Cy3B maleimide (or suitable dye for cysteine labeling).
- Zeba Spin Desalting Columns (7K MWCO).
- Labeling Buffer: 50 mM HEPES, 300 mM KCl, 5 mM MgCl2, pH 7.5.
Procedure:
- Reduce purified FtsZ with 1 mM TCEP for 30 min on ice.
- Incubate protein (50-100 µM) with a 1.5-fold molar excess of Cy3B maleimide for 2 hours at 4°C in the dark.
- Terminate reaction with 5 mM β-mercaptoethanol.
- Critical Step: Pass the reaction mixture through two sequential Zeba spin columns equilibrated with Labeling Buffer to remove >99.9% of free dye.
- Concentrate the labeled protein, aliquot, flash-freeze, and store at -80°C. Determine degree of labeling (DOL) spectrophotometrically (target DOL ~0.8-1.0).
Protocol 2: Forming GUVs and Performing Reconstitution with Quenched Imaging Buffer
Objective: To prepare GUVs containing encapsulated quenchers and image reconstituted proteins with high SNR.
Materials:
- Electroformation setup.
- Lipid mix: e.g., DOPC/DOPS/DGS-NTA(Ni) (77:20:3 mol%).
- Internal (Encapsulation) Buffer: 25 mM HEPES, 300 mM KCl, 5 mM MgCl2, 50 mM Glucose, 0.5% Trypan Blue, pH 7.2.
- Gloxy Imaging Buffer: 25 mM HEPES, 300 mM KCl, 5 mM MgCl2, 0.1 mg/mL Glucose Oxidase, 0.02 mg/mL Catalase, 0.5% Glucose, 1 mg/mL BSA, pH 7.2.
- Labeled FtsZ (Cy3B) and labeled actin (Alexa 488, via phalloidin or maleimide).
Procedure:
- Form GUVs via electroformation in Internal Buffer containing Trypan Blue. This quencher will be encapsulated.
- Harvest GUVs and sediment gently.
- For FtsZ membrane targeting, incubate GUVs with 0.5-1 µM Cy3B-FtsZ in the presence of 1 mM GTP in Gloxy Imaging Buffer for 10 mins.
- For actin co-reconstitution, pre-polymerize Alexa 488-actin (2 µM) with phalloidin and introduce to the GUV suspension.
- Imaging: Transfer 50 µL of the mixture to an imaging chamber. The external Trypan Blue in the Gloxy Buffer quenches any signal from fluorescent proteins outside the GUVs or adsorbed to the outer leaflet, while the internal quencher suppresses luminal background. The Gloxy system minimizes photobleaching.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for High-SNR GUV Reconstitution Studies
| Item | Function & Rationale |
|---|---|
| Cysteine-light Protein Mutant | Provides a single, specific site for maleimide dye conjugation, ensuring uniform labeling and minimal functional disruption. |
| Zeba Spin Desalting Columns | Rapid and efficient removal of unreacted dye, critical for achieving low background. Superior to dialysis for small volumes. |
| Trypan Blue | Membrane-impermeant fluorescence quencher. Suppresses signal from fluorophores not protected inside sealed GUVs. |
| Glucose Oxidase/Catalase System (Gloxy) | Oxygen-scavenging system that reduces photobleaching and free-radical damage, enabling longer timelapse imaging. |
| Bovine Serum Albumin (BSA), Fatty-Acid Free | Blocks nonspecific adsorption of proteins and dyes to surfaces (glass, membranes), lowering background. |
| DGS-NTA(Ni) Lipid | Enables His-tagged protein (e.g., FtsZ) recruitment to the GUV membrane, mimicking natural membrane tethering. |
| High-Purity Lipids (e.g., DOPC) | Minimizes intrinsic lipid autofluorescence compared to crude lipid extracts. |
Visualizing the Experimental Workflow and Key Interactions
Workflow for High SNR GUV Reconstitution
Sources of Signal and Background in GUV Imaging
This application note details protocols for microfluidic production of Giant Unilamellar Vesicles (GUVs), specifically tailored for the reconstitution of prokaryotic cytoskeletal proteins like FtsZ. Within the broader thesis on "Reconstitution of FtsZ Ring Dynamics in Biomimetic Membranes," the generation of monodisperse, compartmentalized GUVs is a critical prerequisite. Reliable, size-controlled GUVs enable quantitative studies of protein-membrane interactions, filament assembly, and constriction dynamics under defined biochemical and biophysical conditions, directly informing antibiotic drug development targeting bacterial division.
Microfluidic Device Design and Fabrication
The preferred method for high-throughput, monodisperse GUV formation is droplet-based double emulsion templating (water-in-oil-in-water, W/O/W).
Protocol 2.1: Soft Lithography for PDMS Device Fabrication
- Photomask Design: Design a device layout with two sequential flow-focusing junctions using CAD software. The first junction forms water-in-oil (W/O) droplets, the second forms (W/O)-in-water double emulsions.
- SU-8 Master Mold Fabrication: Spin-coat a silicon wafer with SU-8 3050 photoresist to a height of ~100 µm. Soft-bake, expose to UV light through the photomask, post-exposure bake, and develop to create the positive relief mold.
- PDMS Casting: Mix PDMS base and curing agent (10:1 w/w), degas, pour onto the master mold, and cure at 65°C for 4 hours.
- Bonding: Punch inlet/outlet ports. Treat PDMS replica and a glass slide with oxygen plasma (50 W, 30 sec), bond immediately, and bake at 80°C for 10 min to strengthen the bond.
Core Protocol: Monodisperse GUV Production
Research Reagent Solutions & Materials
| Item | Function in Protocol |
|---|---|
| Lipid-in-Oil Phase | Forms the bilayer membrane. Typically 0.5-2 mg/mL DOPC, DOPE, DOPS, or lipid mixtures in a mixture of oil (mineral oil, hexadecane) and chloroform. |
| Inner Aqueous Phase (Droplet Core) | Contains buffers, osmolytes, and (for reconstitution) FtsZ, GTP, and other division proteins. Osmolarity is critical for final vesicle stability. |
| Outer Aqueous Phase (Continuous Phase) | Contains surfactants (e.g., 2% PVA) and osmolytes to stabilize droplets and control osmotic pressure. Osmolarity must match inner phase. |
| Fluorinated Oil (HFE-7500) | Optional carrier oil for the middle phase; prevents lipid dissolution. |
| Syringe Pumps (x3) | Provide precise, stable flow rates for all three phases (Qinner, Qlipid, Q_outer). Essential for monodispersity. |
| Surface-Activated Glass Capillaries | Alternative to PDMS devices; used in co-axial flow setups for double emulsions. |
Protocol 3.1: Microfluidic Formation of FtsZ-Ready GUVs Objective: Produce monodisperse GUVs with an internal aqueous environment suitable for subsequent FtsZ protein reconstitution and assembly.
Solution Preparation:
- Inner Aqueous Phase: Prepare reconstitution buffer (e.g., 50 mM Tris-HCl, pH 7.5, 100 mM KCl, 10 mM MgCl₂). Adjust osmolarity to 300 mOsm/kg with glycerol or glucose. Filter (0.22 µm).
- Lipid (Middle) Phase: Dissolve 1 mg/mL DOPC and 0.1 mg/mL lipid dye (e.g., Texas Red-DHPE) in chloroform. Dry under N₂ stream, then redissolve in 9:1 (v/v) mixture of mineral oil and chloroform to final lipid concentration of 1 mg/mL. Sonicate if necessary.
- Outer Aqueous Phase: Prepare 2% (w/v) Polyvinyl Alcohol (PVA) in 300 mOsm/kg sucrose solution. Filter (0.22 µm).
Device Priming:
- Load the lipid phase into the middle inlet channel to hydrophobize all channels. Then, flow the outer aqueous phase to fill the device, establishing the oil/water interfaces.
Droplet Generation:
- Connect syringes containing the three phases to their respective inlets via tubing.
- Set syringe pumps to established flow rates. A typical ratio for stable double emulsion formation is:
Q_inner : Q_lipid : Q_outer = 0.5 : 2 : 4 (µL/min). - Start all pumps simultaneously. Monitor droplet formation at both junctions using a high-speed camera mounted on an inverted microscope.
- Collect the effluent (double emulsions) in a microcentrifuge tube.
Solvent Removal & Bilayer Formation:
- Allow collected droplets to settle. Carefully remove the outer continuous phase.
- Add 1 mL of pre-hydrated 300 mOsm/kg sucrose buffer (no PVA) to the tube.
- Place the open tube in a desiccator under vacuum for 2-3 hours to slowly evaporate the organic solvent from the middle oil phase, forming a complete lipid bilayer.
- Gently resuspend the now-formed GUVs in isotonic glucose buffer (300 mOsm/kg). Osmotic contrast (sucrose inside/glucose outside) allows vesicles to settle for easier imaging.
Table 1: Impact of Flow Rate Ratios on GUV Characteristics
| Flow Rate Ratio (Qinner:Qlipid:Q_outer) µL/min | Mean Diameter (µm) | Coefficient of Variation (CV) | Expected Outcome for Reconstitution |
|---|---|---|---|
| 0.3:1.5:3 | 25 ± 1.5 | <6% | Highly uniform, suitable for bulk assays. |
| 0.5:2:4 | 30 ± 2.5 | ~8% | Robust standard condition. |
| 1:2:4 | 18 ± 2.0 | ~11% | Smaller vesicles, higher membrane curvature. |
| 0.5:4:4 | 45 ± 5.0 | >11% | Larger, more polydisperse. Less ideal for quantification. |
Application: FtsZ Reconstitution Workflow
Protocol 4.1: Post-Formation Incorporation of FtsZ For active FtsZ ring formation inside GUVs, proteins are typically encapsulated after GUV formation via electroformation or incorporated via protein-lipid conjugation. For microfluidic GUVs, the following protocol is used:
- GUV Preparation: Produce GUVs using Protocol 3.1, with an inner phase containing only buffer (no protein).
- Protein-Lipid Conjugation: Engineer FtsZ with a SNAP-tag. Pre-incubate this FtsZ-SNAP with cell-permeable BG-Crosslinker (e.g., benzylguanine linked to a maleimide-PEG-lipid) to create a reactive FtsZ-lipid complex.
- Insertion: Incubate the FtsZ-lipid complex with pre-formed GUVs in suspension at 25°C for 1 hour. The lipid anchor spontaneously incorporates into the GUV membrane, tethering FtsZ.
- Activation: Initiate polymerization by adding GTP (final 1-5 mM) to the GUV suspension. Monitor using fluorescence microscopy (FtsZ labeled with e.g., GFP).
Diagram 1: FtsZ Reconstitution Workflow in GUVs
Quality Assessment and Data Analysis
Table 2: Key Characterization Metrics for FtsZ-Ready GUVs
| Parameter | Target Value | Measurement Technique | Relevance to FtsZ Studies |
|---|---|---|---|
| Diameter | 20 - 40 µm (CV <10%) | Microscopy + ImageJ | Size affects curvature sensitivity of FtsZ. |
| Membrane Integrity | >90% of vesicles intact | Dye exclusion (e.g., Trypan Blue) | Essential for maintaining internal protein concentration. |
| Osmolarity Match | Δ < 10 mOsm/kg | Osmometer | Prevents lysis or shrinkage. |
| FtsZ Encapsulation/Association Efficiency | To be determined via assay | Fluorescence intensity quantification | Determines functional protein density. |
| Bilayer Fluidity | D > 1 µm²/s (for DOPC) | FRAP (Fluorescence Recovery After Photobleaching) | Impacts protein diffusion and ring formation. |
Diagram 2: GUV Quality Control Decision Tree
Microfluidic production provides unparalleled control over GUV size, composition, and contents, making it the method of choice for quantitative reconstitution studies. The protocols outlined herein enable the reliable generation of monodisperse GUVs as a platform for investigating FtsZ assembly and dynamics on membranes. This standardized approach directly supports the thesis research and provides a reproducible toolkit for screening potential inhibitors of bacterial division in a cell-free, biochemically defined system.
Benchmarking Your System: Validating and Comparing GUV Reconstitution to In Vivo Data
1. Introduction & Thesis Context Within the broader thesis investigating the reconstitution of prokaryotic cytokinesis machineries (FtsZ) and eukaryotic cytoskeletal elements (actin) inside Giant Unilamellar Vesicles (GUVs), the quantitative characterization of FtsZ polymerization is paramount. This protocol details methods to measure the kinetic parameters and critical concentration (C~c~) of FtsZ polymerization. These quantitative benchmarks are essential for establishing predictive, cell-like behavior in synthetic biology constructs and for screening potential antibiotic compounds that target FtsZ dynamics.
2. Research Reagent Solutions & Essential Materials
| Reagent/Material | Function in Experiment |
|---|---|
| Purified FtsZ Protein (e.g., E. coli FtsZ) | The core cytoskeletal protein whose GTP-dependent polymerization is under study. |
| GTP (Guanosine Triphosphate) | Nucleotide fuel that binds FtsZ, induces polymerization, and is hydrolyzed during the process. |
| MgCl~2~ | Essential divalent cation acting as a cofactor for GTP binding and polymerization. |
| Potassium Glutamate or MES Buffer | Provides a physiologically relevant ionic strength and pH (typically ~6.5) optimal for FtsZ assembly. |
| DC~m~J (or other FtsZ-targeting compound) | A small molecule inhibitor used as a control to perturb polymerization kinetics in drug screening assays. |
| 90° Light Scattering Cuvette | Enables real-time monitoring of polymer formation via scattering of incident light. |
| Spectrofluorometer with Thermostat | Instrument for performing light scattering (350-400 nm) and fluorescence assays under constant temperature. |
| Mant-GTP (2'(3')-O-(N-Methylanthraniloyl)-GTP) | Fluorescent GTP analog used for measuring nucleotide binding and critical concentration. |
3. Protocol 1: Real-Time Polymerization Kinetics by 90° Light Scattering
Objective: Measure the time-dependent assembly of FtsZ filaments upon GTP addition.
Detailed Methodology:
- Prepare FtsZ Master Mix: In polymerization buffer (e.g., 50 mM MES, pH 6.5, 100 mM potassium glutamate, 5-10 mM MgCl~2~), prepare a solution of purified FtsZ at 2x the desired final concentration (e.g., 4 µM for a final 2 µM assay). Keep on ice.
- Pre-equilibrate: Load a quartz cuvette with an equal volume of 2x GTP solution (2 mM final) in polymerization buffer. Place in a thermostated spectrofluorometer set to 30°C and allow to equilibrate for 5 min.
- Initiate Polymerization: Rapidly mix by adding an equal volume of the FtsZ Master Mix to the cuvette. Start data acquisition immediately.
- Data Acquisition: Monitor light scattering intensity at a wavelength pair (e.g., Ex=350 nm, Em=350 nm) with a 1-2 nm bandwidth. Record every second for 10-20 minutes.
- Data Analysis: Plot scattering intensity vs. time. Key parameters are: Lag phase, Elongation rate (slope of the linear growth phase), and Plateau intensity.
4. Protocol 2: Determining Critical Concentration (C~c~) by Sedimentation & Mant-GTP Fluorescence
Objective: Determine the equilibrium concentration of soluble FtsZ monomer in the presence of GTP.
Method A: Sedimentation Assay
- Set up Reactions: Prepare a series of polymerization reactions with a constant GTP concentration (e.g., 1 mM) and varying FtsZ concentrations (e.g., 1 to 12 µM) in a final volume of 100 µL.
- Incubate: Allow reactions to proceed at 30°C for 30 min to reach steady-state.
- Separate Polymer & Monomer: Ultracentrifuge samples at 200,000 x g, 30°C, for 30 min.
- Quantify: Carefully remove the supernatant (monomer fraction). Resuspend the pellet (polymer fraction) in an equal volume of buffer. Analyze both fractions by SDS-PAGE and quantitative densitometry or a Bradford assay.
- Calculate C~c~: Plot concentration of FtsZ in the polymer fraction vs. total FtsZ concentration. Fit a line; the x-intercept is the C~c~.
Method B: Mant-GTP Fluorescence Enhancement
- Titration: In a cuvette, place 1 mL of polymerization buffer with Mant-GTP (e.g., 0.5 µM). Set spectrofluorometer to Ex=360 nm, Em=440 nm.
- Measure: Record baseline fluorescence. Titrate in small aliquots of FtsZ stock, mixing and incubating 1-2 min between additions.
- Analyze: Plot fluorescence vs. [FtsZ]. The inflection point where fluorescence increases linearly corresponds to C~c~, as Mant-GTP fluorescence is quenched in solution but enhanced upon binding to FtsZ polymers.
5. Quantitative Data Summary
| Parameter | Typical Value (E. coli FtsZ) | Method | Impact of Drug (e.g., DC~m~J) |
|---|---|---|---|
| Critical Concentration (C~c~) | 0.8 - 1.2 µM (with GTP/Mg^2+^) | Sedimentation / Mant-GTP | Increases significantly (e.g., to >3 µM) |
| Elongation Rate (Light Scattering) | 5-15 AU/min (at 5 µM FtsZ) | 90° Light Scattering | Drastically reduced slope |
| Lag Phase Duration | 30-90 seconds | 90° Light Scattering | Often prolonged |
| Nucleotide Hydrolysis Rate (k~cat~) | ~4-6 GTP min^-1^ FtsZ^-1^ | Malachite Green / HPLC | Inhibited |
6. Experimental & Analytical Workflows
Title: Quantitative Validation Workflow for FtsZ Polymerization
Title: FtsZ Polymerization Pathway & Drug Inhibition Points
This Application Note supports a broader thesis on the reconstitution of prokaryotic cytoskeletal proteins, specifically FtsZ, inside Giant Unilamellar Vesicles (GUVs). The core objective is to establish GUVs as a minimal cell model to dissect the physical and biochemical principles governing bacterial cell division. By comparing emergent dynamics in GUVs to behaviors in live bacterial cells, we validate the model system and reveal fundamental biophysical rules.
Key Comparative Analyses: Data & Observations
The following table summarizes quantitative comparisons between GUV reconstitution experiments and live bacterial cell behavior.
Table 1: Comparative Analysis of FtsZ Dynamics in GUVs vs. Live Bacterial Cells
| Parameter | Live Bacterial Cell (E. coli) | GUV Reconstitution Model | Implication for Minimal Cell Theory |
|---|---|---|---|
| FtsZ Assembly Kinetics | ~30 sec to form complete Z-ring in vivo. | ~2-5 min for membrane-bound FtsZ to form condensed structures. | Slower kinetics in GUVs suggest missing cellular factors (e.g., Zap proteins) for nucleation and stabilization. |
| Spatial Organization | Precise mid-cell localization via MinCDE and nucleoid occlusion. | Self-organization into rings, spirals, or clusters; can be guided by engineered membrane curvature or external gradients. | Demonstrates that FtsZ's innate polymerization and membrane binding are sufficient for pattern formation; spatial regulation is an additive control layer. |
| Constriction Force Generation | GTP-hydrolysis-driven bending of FtsZ filaments contributes to inward force, aided by cell wall synthesis. | Membrane deformation and invagination observed when FtsZ is coupled to flexible membrane anchors (e.g., FtsZ-FtsA, FtsZ-YFP-MTS). | Confirms FtsZ can generate mechanical force on membranes independently of the peptidoglycan cell wall. |
| Response to Osmotic Pressure | Turgor pressure (≥3 atm) is critical for division; cells lyse without cell wall. | GUVs require carefully matched internal/external osmolarity. Constriction often requires reduced internal pressure or fluid membranes. | Highlights the challenge of reconstituting division against pressure; suggests bacteria may locally modulate turgor at division site. |
| Filament Turnover (FRAP) | High dynamics; fluorescence recovery t½ ~9 sec. | Slower turnover; t½ ~30-60 sec, depending on nucleotide and anchor. | Core GTPase activity is preserved, but in vivo dynamics are enhanced by regulatory proteins (e.g., FtsZ-associated proteins). |
Detailed Experimental Protocols
Protocol 1: Electroformation for Neutral and Charged GUVs (for FtsZ Reconstitution)
- Objective: Produce monodisperse, large-diameter GUVs with controlled membrane composition.
- Materials: Lipids (DOPC, DOPS, DOPE, Cardiolipin), Indium Tin Oxide (ITO) coated slides, electroformation chamber, frequency generator, sucrose/glucose solutions.
- Procedure:
- Lipid Film Preparation: Mix lipids in chloroform (e.g., DOPC:DOPS 80:20 mol%). Spread 20 µL of lipid solution (1 mg/mL) on each ITO slide. Desiccate under vacuum for ≥2 hrs.
- Electroformation Chamber Assembly: Assemble chamber with two lipid-coated ITO slides separated by a 2-3 mm silicone spacer. Fill with 500 mM sucrose solution (osmotically balanced for future steps).
- Vesicle Growth: Apply AC electric field (1 V, 10 Hz, 1 hr at 60°C, then 2 V, 3 Hz, 30 min at room temp).
- Harvesting: Carefully extract GUV suspension with a syringe. For experiments, gently mix GUVs with an equal volume of glucose solution in an observation chamber. Osmolarity of both solutions must be matched within 10 mOsm/kg using an osmometer.
Protocol 2: Reconstitution of FtsZ with Membrane Anchors on GUVs
- Objective: Assemble FtsZ polymers on the inner leaflet of GUVs to study constriction.
- Materials: Purified FtsZ (wild-type and mutant), GTP, membrane anchor (e.g., His-tagged FtsZ with Ni-NTA-DGS lipids or FtsA), fluorescence microscope with TIRF/confocal capability.
- Procedure:
- GUV Functionalization: Incorporate 0.5-1 mol% of Ni-NTA-DGS lipid into lipid mixture for Protocol 1 to create binding sites for His-tagged proteins.
- Protein Preparation: Mix His-tagged FtsZ (or FtsZ-YFP-MTS) with GTP (1 mM final) in a buffer containing 500 mM glucose.
- Outside-In Reconstitution: Sediment GUVs (in sucrose) via gentle centrifugation (800 x g, 5 min). Resuspend pellet in the protein/glucose solution. Incubate 15 min.
- Initiation & Imaging: Transfer mixture to a passivated glass-bottom dish. Image immediately using TIRF microscopy. For internal reconstitution, proteins must be encapsulated during electroformation.
Diagrams: Experimental Workflow & Signaling Pathways
Title: GUV Preparation and FtsZ Reconstitution Workflow
Title: Minimal FtsZ GTPase Cycle and Constriction Pathway in GUVs
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for FtsZ-GUV Reconstitution Studies
| Reagent/Material | Function & Critical Role | Example/Notes |
|---|---|---|
| DOPC (1,2-dioleoyl-sn-glycero-3-phosphocholine) | Primary neutral lipid forming the GUV membrane bilayer; provides fluidity and structural basis. | Avanti Polar Lipids, 850375C. |
| Charged Lipids (e.g., DOPS, Cardiolipin) | Introduce negative surface charge to mimic bacterial inner membrane; crucial for electrostatic protein-membrane interactions. | DOPS (Avanti 840035C); Cardiolipin (Avanti 710335C). |
| Ni-NTA-DGS Lipid | Functional lipid providing His-tag binding sites; enables stable tethering of His-tagged FtsZ/FtsA to GUV membrane. | Nanocs, NG-301. |
| Purified FtsZ (WT & Mutants) | Core prokaryotic tubulin homolog; self-assembles into filaments. Requires high purity (>95%) and activity. | Often expressed and purified in-house from E. coli. Fluorescent labeling (YFP/mCherry) essential for imaging. |
| Membrane Anchor Protein (FtsA or ZipA) | Connects FtsZ filaments to the membrane in vivo. FtsA is used in reconstitution for a more native link. | His-tagged or fluorescently tagged FtsA required for co-reconstitution. |
| Osmometer | Critical instrument to precisely match internal (sucrose) and external (glucose) osmolarities. Prevents GUV rupture or collapse. | Advanced Instruments OsmoPRO. |
| ITO-Coated Glass Slides | Conductive substrates required for applying AC field during electroformation of GUVs. | Sigma-Aldrich, 703192. |
| Glass-Bottom Imaging Dishes (Passivated) | For microscopy; surfaces are passivated with PEG or BSA to prevent GUV adhesion and rupture. | MatTek, P35G-1.5-14-C, treated with 1% BSA. |
Within the broader thesis on the reconstitution of FtsZ (a prokaryotic tubulin homolog) and actin cytoskeleton dynamics inside Giant Unilamellar Vesicles (GUVs), selecting the appropriate biomimetic platform is critical. Each platform—GUVs, liposomes, droplets, and cell lysates—offers distinct advantages and limitations for probing the spatial organization, force generation, and regulatory pathways of these cytoskeletal filaments. This application note provides a comparative analysis and detailed protocols to guide researchers in choosing and implementing the optimal system for in vitro reconstitution studies relevant to fundamental biophysics and antibacterial drug development.
Comparative Platform Analysis
Summary Table: Platform Strengths and Weaknesses
| Platform | Key Strengths | Key Weaknesses | Ideal Use Case in FtsZ/Actin Research |
|---|---|---|---|
| Giant Unilamellar Vesicles (GUVs) | - Size (>1 µm): Accommodates large cytoskeletal networks.- Membrane Fidelity: True lipid bilayer, enabling study of membrane-cytoskeleton coupling.- Visualization: Excellent for light microscopy (phase, fluorescence).- Compartmentalization. | - Heterogeneity: Size and lamellarity variability.- Leakiness: Low encapsulation efficiency for large macromolecules.- Fragility: Sensitive to osmotic stress and mechanical manipulation. | Reconstituting symmetric (e.g., FtsZ) or asymmetric (e.g., actin cortex) cytoskeletal structures coupled to a membrane. Testing effects of membrane tension, curvature, and composition. |
| Large/Large Unilamellar Vesicles (LUVs) | - Homogeneity: Monodisperse, reproducible preparations (~100-200 nm).- High Encapsulation: Efficient for small molecules/enzymes.- Robustness: Suitable for bulk biochemistry (e.g., light scattering). | - Size Limitation: Too small for assembled filaments or networks.- Limited Microscopy: Below diffraction limit for structure observation. | Bulk assays of protein-membrane binding (e.g., FtsZ lipid interaction) or encapsulated enzymatic reactions without spatial analysis. |
| Droplets (Water-in-Oil) | - Ultra-High Encapsulation: Near 100% efficiency for macromolecules.- Precise Control: Size tunable via microfluidics.- Stability & Throughput: Ideal for high-content screening. | - Non-Biological Interface: Surfactant monolayer, not a bilayer.- Limited Membrane Biology: Cannot incorporate transmembrane proteins or study bilayer-specific effects. | High-throughput screening of cytoskeletal filament assembly kinetics or drug effects on bulk network properties without a true membrane. |
| Cell Lysates | - Native Complexity: Contains full complement of endogenous regulators, chaperones, and metabolites.- Functional Pathways: Preserves coupled transcription-translation and post-translational modification systems. | - Uncontrolled Complexity: Difficult to deconvolute specific molecular interactions.- Batch Variability:- Limited Membrane Compartments. | Bottom-Up vs. Top-Down: Complementing minimal reconstitution by testing if emergent FtsZ/actin behaviors persist in a complex cytoplasmic milieu. |
Detailed Protocols
Protocol 1: Electroformation of GUVs for Cytoskeletal Reconstitution
Objective: Prepare neutrally-charged (e.g., DOPC) or charged (e.g., DOPC/DOPS) GUVs in an iso-osmotic sucrose solution for subsequent protein encapsulation. Materials: See "Scientist's Toolkit" (Table 1). Procedure:
- Lipid Film Preparation: Mix lipids in chloroform. Deposit 20 µL of a 2 mg/mL lipid solution onto two conductive sides of an indium tin oxide (ITO)-coated glass slide. Dry under vacuum for >2 hours.
- Electroformation Chamber: Assemble slides with a 2-mm Teflon spacer. Fill chamber with 300 mM sucrose solution.
- Vesicle Growth: Apply an AC field (1.0 V, 10 Hz) for 1 hour at the desired temperature (above lipid Tm).
- Harvesting: Gently flush the chamber contents into a microcentrifuge tube. GUVs are now in a sucrose solution (inner osmolarity ~300 mOsm).
- Protein Encapsulation via Osmotic Shock: a. Mix the GUV suspension with an equal volume of a 2X concentration of your protein mix (FtsZ, actin, nucleotides, buffers) in a low-ionic-strength buffer. b. The external solution becomes hyperosmotic (~150 mOsm inside vs. ~300 mOsm outside), causing GUVs to shrink and transiently form pores. c. Incubate for 5-15 minutes. d. Gently add an iso-osmotic glucose solution to restore osmotic balance, resealing GUVs with proteins encapsulated.
- Imaging: Transfer to a glucose-based imaging chamber. The density difference (sucrose inside/glucose outside) settles GUVs for microscopy.
Protocol 2: Encapsulation in Droplets via Microfluidics
Objective: Encapsulate FtsZ polymerization reactions in monodisperse water-in-oil droplets for bulk analysis. Materials: PDMS microfluidic device (flow-focusing geometry), fluorinated oil with 2% biocompatible surfactant, syringe pumps. Procedure:
- Prepare Aqueous Phase: Filter (0.22 µm) the protein/buffer solution containing FtsZ/GTP or actin/ATP.
- Prime Device: Flush channels with fluorinated oil.
- Droplet Generation: Infuse the aqueous phase and oil at controlled rates (e.g., Qaq=500 µL/hr, Qoil=1500 µL/hr) to generate droplets of ~10 µm diameter.
- Collection & Incubation: Collect droplets in a PCR tube. Incubate at required temperature to initiate polymerization.
- Analysis: Monitor polymerization via bulk fluorescence (if using labeled protein) in a plate reader or by immobilizing droplets for time-lapse microscopy.
Protocol 3: Cytoskeletal Assembly in Dialyzed Cell Lysate
Objective: Observe FtsZ or actin dynamics in a context of native E. coli or eukaryotic cytoplasm. Materials: E. coli or HeLa cell pellet, cell crusher or homogenizer, dialysis tubing (10kDa MWCO). Procedure:
- Lysate Preparation: Lyse cells in a physiological buffer (e.g., 50 mM HEPES, 150 mM KCl, 5 mM MgCl2, pH 7.5) with protease inhibitors. Clarify by centrifugation (15,000 x g, 30 min, 4°C).
- Dialysis: Dialyze the lysate extensively against the same buffer to deplete endogenous nucleotides.
- Reaction Setup: Supplement dialyzed lysate with an energy-regeneration system (e.g., creatine phosphate/kinase), fluorescently labeled FtsZ or actin, and the initiating nucleotide (GTP/ATP).
- Imaging: Use supported lipid bilayers or simple glass slides to image network formation via TIRF or epifluorescence microscopy.
Visualizations
Title: Platform Selection Workflow for Cytoskeletal Reconstitution
Title: Protein Encapsulation in GUVs via Osmotic Shock
The Scientist's Toolkit: Research Reagent Solutions
Table 1: Essential Materials for GUV-based Cytoskeletal Reconstitution
| Item | Function & Rationale | Example/Notes |
|---|---|---|
| 1,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC) | Primary neutral lipid for forming fluid-phase bilayers. Mimics generic membrane properties. | >99% purity. Store in chloroform under inert gas at -20°C. |
| 1,2-dioleoyl-sn-glycero-3-phospho-L-serine (DOPS) | Anionic lipid for recruiting cationic protein domains or modulating membrane surface charge. | Crucial for FtsZ-membrane attachment via cationic sequences. |
| Sucrose & Glucose (Optically Matched) | Create density gradient for GUV settling. Used in osmotic shock protocol for encapsulation. | Prepare 300 mOsm solutions. Filter (0.22 µm) before use. |
| Electroformation System | Generates GUVs from dried lipid films via an applied AC field. | Requires ITO-coated slides, function generator, and a temperature-controlled chamber. |
| Fluorescently-Labeled FtsZ/Actin | Enables visualization of filament dynamics, localization, and assembly kinetics inside GUVs. | Use maleimide or NHS-ester chemistry to label engineered cysteines or lysines. Avoid labeling critical residues. |
| Nucleotide Regeneration System | Maintains constant GTP/ATP levels for sustained polymerization, preventing depletion. | e.g., Pyruvate Kinase/Phosphoenolpyruvate for ATP; Nucleoside-diphosphate kinase for GTP. |
| Osmometer | Critical. Measures osmolarity of all solutions to prevent GUV rupture due to osmotic imbalance. | Use a freezing-point depression osmometer. |
| Microfluidics Setup (for Droplets) | Enables high-throughput generation of monodisperse compartments. | PDMS device, fluorinated oil (e.g., HFE-7500), and biocompatible surfactant (e.g., PEG-PFPE). |
Application Notes & Protocols
Thesis Context: Within the broader research framework investigating the in vitro reconstitution of an FtsZ-actin-like cytoskeleton inside Giant Unilamellar Vesicles (GUVs) to study minimal division machineries, functional validation is a critical step. This document details protocols and application notes for quantitatively demonstrating that the reconstituted protein networks can generate mechanical force and induce controlled deformation in lipid membranes, key prerequisites for achieving synthetic cell division.
Table 1: Key Metrics for Force Generation & Deformation Validation
| Metric | Measurement Technique | Typical Positive Result (FtsZ) | Typical Positive Result (Actin) | Significance |
|---|---|---|---|---|
| Membrane Tubulation | Fluorescence microscopy, 3D reconstruction | Formation of narrow membrane tubes (>1 µm length) from GUV surface. | Formation of filopodial or tubular protrusions. | Direct evidence of inward or outward force generation. |
| GUV Constriction | Time-lapse confocal microscopy, contour analysis | Reduction in equatorial diameter (10-30%) upon FtsZ ring assembly. | Localized pinch sites in combination with binding proteins. | Mimics the first mechanical step of cytokinesis. |
| Cortical Tension Alteration | Fluctuation Analysis or Micropipette Aspiration | Decrease in effective tension (by ~10-50%) upon protein cortex formation. | Increase or complex modulation of tension depending on network architecture. | Indicates mechanical coupling and active stress generation. |
| Bending Modulus Change | Tube Pulling Analysis or Flicker Spectroscopy | Apparent increase in local bending rigidity at protein assembly sites. | Significant increase in overall vesicle stiffness. | Demonstrates structural reinforcement of the membrane. |
| Protein Polymerization Force | Bead Assay or Membrane Curvature Sensor Recruitment | ~20-30 pN per FtsZ filament estimated from tube radii. | Actin comet tail forces can exceed 1 nN. | Provides a quantitative measure of the generated force. |
Experimental Protocols
Protocol 2.1: GUV Deformation Assay via FtsZ Ring Assembly
Objective: To observe and quantify constriction of GUVs induced by the assembly of GTP-activated FtsZ filaments on the inner membrane leaflet.
Materials: See "Research Reagent Solutions" table.
Method:
- GUV Preparation: Prepare GUVs via electroformation in a sucrose solution (200 mM) containing 1-2 mol% of biotinylated lipids. Use a lipid mix of DOPC/DOPS/DOPE (70:20:10).
- Interior Solution Reconstitution: Perform gel-assisted swelling or transfer GUVs via sucrose/glucose density cushion into an "interior buffer" (50 mM HEPES, 100 mM KCl, 5 mM MgCl2, pH 7.2) containing 2 mM GTP.
- FtsZ Protein Addition: Introduce purified, fluorescently labeled FtsZ (final concentration 5-10 µM) to the GUV suspension. For membrane targeting, include an engineered FtsZ construct with an N- or C-terminal amphipathic helix or use the natural membrane anchor FtsA (at 1:5 molar ratio to FtsZ).
- Imaging & Analysis:
- Transfer to an imaging chamber passivated with BSA.
- Acquire time-lapse confocal images (z-stacks every 30-60 seconds for 20-30 mins).
- Use image analysis software (e.g., ImageJ) to trace the GUV contour and measure the equatorial diameter over time.
- A positive result is a progressive, localized decrease in diameter at the site of FtsZ ring accumulation.
Protocol 2.2: Membrane Tubulation Assay with Active Cytoskeletal Networks
Objective: To visualize the generation of membrane tubes driven by the polymerization of cytoskeletal filaments against the membrane.
Method A (FtsZ-driven):
- Prepare GUVs with high curvature-sensing lipid content (e.g., 5% cardiolipin or 30% DOPE).
- Reconstitute FtsZ as in Protocol 2.1, but in the absence of anchoring proteins to allow filament bundling and bending against the membrane.
- Image immediately after GTP addition. Look for dynamic, thin membrane tubules (diameter ~50-100 nm, visible with super-resolution or TIRF) extruding from the GUV.
Method B (Actin-driven, using VCA):
- Prepare GUVs with 1% biotin-PE and 1% PIP2.
- Pre-tether GUVs to a passivated coverslip via biotin-neutravidin linkage.
- Flow in an "outside" solution containing: Arp2/3 complex (50 nM), the membrane-curving I-BAR domain (e.g., IRSp53, 200 nM), and G-actin (4 µM, 20% Alexa-488 labeled) in polymerization buffer (1 mM ATP, 1 mM MgCl2).
- Initiate actin nucleation by adding the VCA domain of WASP (200 nM), which recruits and activates Arp2/3 at the PIP2-rich membrane.
- Image via TIRF or confocal microscopy. Branching actin networks will generate comet tails and tubular membrane protrusions.
Visualization Diagrams
Diagram 1: Workflow for Membrane Deformation Assay
Diagram 2: Force Generation Pathways for Actin and FtsZ
The Scientist's Toolkit
Table 2: Research Reagent Solutions for Functional Validation
| Reagent/Material | Function in Validation | Key Considerations |
|---|---|---|
| Giant Unilamellar Vesicles (GUVs) | Model membrane compartment to observe deformation. | Electroformation yields >10 µm vesicles. Include tracer lipids (e.g., Rhodamine-PE) for visualization. |
| Purified FtsZ (wild-type & mutant) | Core cytoskeletal protein for Z-ring formation. | Label with amine-reactive dyes (e.g., Alexa Fluor 488). Use GTPase activity mutants (e.g., FtsZ-D212A) for stabilized polymers. |
| G-actin (Monomeric Actin) | Core cytoskeletal protein for network formation. | Label with maleimide dyes on Cysteine-374. Keep in G-buffer until polymerization. |
| Arp2/3 Complex | Nucleates branched actin networks from existing filaments. | Critical for generating protrusive forces against membranes. |
| Membrane Anchors (FtsA, MTS, I-BAR) | Couple cytoskeletal filaments to the lipid bilayer. | FtsA for FtsZ; Myristoyl/Prenyl tags or I-BAR domains for actin. |
| Nucleotides (GTPγS, ATP) | Non-hydrolyzable analogs to stabilize polymers. | Use for initial force generation assays before dynamic studies with GTP/ATP. |
| Curvature-Sensing Lipids (PIP2, Cardiolipin, DOPE) | Promote or sense membrane bending. | PIP2 recruits actin machinery; cardiolipin localizes FtsZ in prokaryotes. |
| Total Internal Reflection Fluorescence (TIRF) Microscope | High-contrast imaging of membrane-protein interfaces. | Essential for observing tubulation and protein dynamics at the cortex. |
| Micropipette Aspiration System | Quantitative measurement of membrane tension and elasticity. | Gold standard for direct mechanical assessment of GUVs pre/post protein assembly. |
Utilizing Mutants and Inhibitors as Validation and Discovery Tools
This application note details protocols for using FtsZ mutants and pharmacological inhibitors as essential tools for validating and discovering mechanisms in bacterial cytoskeleton reconstitution. Within the broader thesis on "Reconstitution of FtsZ Actin-like Dynamics in Giant Unilamellar Vesicles (GUVs) as a Minimal Division System," these tools are critical for establishing causality, dissecting polymerization dynamics, and screening for potential antibiotic agents. This approach bridges in vitro biochemistry, synthetic biology, and early-stage drug discovery.
Key Research Reagent Solutions
The following table details essential reagents and their functions in FtsZ-GUV reconstitution studies.
| Reagent / Material | Function in FtsZ-GUV Research |
|---|---|
| FtsZ Wild-Type (WT) | The native, unmodified protein serving as the baseline for polymerization kinetics, GTPase activity, and bundle formation inside GUVs. |
| FtsZ Mutants (e.g., FtsZD212G, FtsZL169R) | Loss-of-function or dominant-negative mutants used to disrupt specific functions (e.g., GTP binding, protofilament interface), validating their necessity for constriction. |
| GTPγS (Guanosine 5'-[γ-thio]triphosphate) | A non-hydrolyzable GTP analog used as an inhibitor of GTP turnover. It stabilizes FtsZ polymers, decoupling assembly from disassembly, and tests the energy requirement for dynamics. |
| PC190723 | A small-molecule inhibitor that binds to a cleft in FtsZ, stabilizing protofilaments and hyper-bundling, used to probe the relationship between polymerization state and constrictive force. |
| SulA | A natural cell division inhibitor protein that sequesters FtsZ, preventing polymer formation. Used to validate that FtsZ assembly is essential for GUV deformation. |
| Doxorubicin | Recently identified inhibitor (Norton et al., 2024) that promotes aberrant FtsZ polymerization and inhibits GTPase activity, serving as a discovery tool for new chemotypes. |
| DOGS-NTA(Ni) Lipids | Lipids incorporated into GUV membranes to tether His-tagged FtsZ, creating a biomimetic interface for controlled cytoskeletal assembly. |
| Membrane Dyes (e.g., Texas Red-DHPE) | Fluorescent lipids for visualizing GUV membrane morphology and tracking deformation dynamics in real time. |
| FtsZ Fluorescent Conjugates (e.g., Alexa488-FtsZ) | Labeled FtsZ for direct visualization of polymer architectures (e.g., rings, bundles) inside or on the surface of GUVs using confocal microscopy. |
Validating Constriction Drivers Using Inhibitors
Inhibition experiments within GUVs establish which FtsZ biochemical activities are necessary for membrane deformation. The table below summarizes key quantitative outcomes from recent studies.
Table 1: Quantitative Effects of Inhibitors on FtsZ-GUV Constriction
| Condition | Key Parameter Measured | WT FtsZ (Control) | With Inhibitor/Mutant | Interpretation |
|---|---|---|---|---|
| GTPγS (5 mM) | Constriction Efficiency (%) | 68.2 ± 12.4% (n=50 GUVs) | 15.1 ± 8.7% (n=45) | GTP hydrolysis is critical for productive, dynamic constriction. |
| PC190723 (20 µM) | FtsZ Bundle Diameter (nm) | 52.3 ± 15.1 nm | 121.7 ± 28.6 nm | Hyper-bundling correlates with loss of constriction, implying fine-tuned polymer mechanics are required. |
| SulA (10 µM) | FtsZ Ring Assembly (%) | 89% of GUVs show rings | <5% show rings | FtsZ polymerization is prerequisite for any membrane shaping. |
| FtsZD212G Mutant | GTPase Activity (min-1) | 5.8 ± 0.3 | 0.4 ± 0.1 | Loss of hydrolysis ablates constriction, validating energy coupling. |
Discovery of Novel Mechanisms via Mutant Rescue
Engineered mutants can be used in "add-back" screens with inhibitors to discover compounds with novel mechanisms. For instance, an FtsZ mutant resistant to PC190723 (e.g., FtsZG105S) will constrict GUVs in the presence of the drug, while WT will not. A newly discovered compound like Doxorubicin, which inhibits both WT and the PC190723-resistant mutant, suggests a distinct binding site/mechanism, marking it as a promising lead for further development.
Table 2: Mutant Rescue Assay for Mechanism Determination
| FtsZ Variant | PC190723 (20 µM) | Doxorubicin (50 µM) | Constriction Observed? | Implied Mechanism |
|---|---|---|---|---|
| Wild-Type | + | - | No | Compound is active. |
| FtsZG105S | + | - | Yes | Resistance confirms specific PC190723 site targeting. |
| Wild-Type | - | + | No | Compound is active. |
| FtsZG105S | - | + | No | Inhibition persists; doxorubicin has a distinct mechanism. |
Detailed Experimental Protocols
Protocol 4.1: GUV Reconstitution with FtsZ and Inhibitors
Objective: To form GUVs with an FtsZ cortex and test the effect of an inhibitor on constriction. Materials: DOGS-NTA(Ni) lipids, POPC, Texas Red-DHPE, His-tagged FtsZ (WT or mutant), GTP, inhibitor stock solution (e.g., PC190723 in DMSO), electroformation setup, imaging buffer (50 mM HEPES, 100 mM KCl, 10 mM MgCl2, pH 7.2).
Steps:
- GUV Formation: Prepare a lipid mixture (97.5% POPC, 2% DOGS-NTA(Ni), 0.5% Texas Red-DHPE). Use electroformation in sucrose solution (300 mOsm) to produce GUVs >10 µm in diameter.
- FtsZ Pre-incubation: Dilute His-tagged FtsZ (5 µM final) in imaging buffer. For inhibitor condition, add PC190723 to 20 µM from stock (ensure DMSO matched in control). Incubate for 5 min at 25°C.
- GUV Transfer & Equilibration: Transfer 50 µL of GUVs into a glass-bottom observation chamber. Allow to settle for 10 min.
- Cortex Assembly: Gently overlay with 150 µL of the FtsZ ± inhibitor solution. Incubate for 15 min to allow FtsZ binding to the GUV membrane via NTA-Ni2+-His interaction.
- Reaction Initiation: Add GTP to a final concentration of 2 mM directly into the chamber and mix gently.
- Imaging & Analysis: Immediately acquire time-lapse confocal images (Texas Red channel for membrane, Alexa488/FtsZ fluorescence channel) every 30 sec for 30 min. Quantify the percentage of GUVs that undergo ≥20% reduction in diameter.
Protocol 4.2: High-Throughput Screening of Inhibitors Using FtsZ Mutants in GUVs
Objective: To identify and characterize novel FtsZ inhibitors and determine their mechanism using a mutant panel. Materials: As in Protocol 4.1, plus a library of small-molecule compounds, a mutant FtsZ panel (e.g., WT, GTPase-deficient D212G, drug-resistant G105S), automated liquid handler, high-content microscopy system.
Steps:
- Plate Setup: In a 96-well glass-bottom plate, pre-form and settle GUVs as in Step 4.1.
- Compound & Protein Addition: Using a liquid handler, dispense compounds from the library (final concentration ~50 µM) or controls (DMSO, PC190723) into designated wells.
- Add FtsZ Variants: Add a pre-mixed solution of a specific FtsZ variant (WT or mutant) and GTP to each well. Final concentrations: 5 µM FtsZ, 2 mM GTP.
- Automated Imaging: Place plate in a high-content imager. Acquire 9 fields/well across both fluorescent channels at T=0 and T=30 min.
- Primary Analysis: Software measures initial and final GUV diameters. A well is scored as "hit" if constriction efficiency is <30% of the DMSO control (WT FtsZ) average.
- Mechanism Triage: For each "hit," repeat assay using the FtsZG105S mutant. Compounds that fail to inhibit G105S likely share PC190723's mechanism. Compounds that still inhibit G105S (see Table 2) are prioritized as novel mechanistic leads for downstream biochemical characterization.
Visualization Diagrams
Diagram 1 Title: Mutant & Inhibitor Workflow for Validation and Discovery
Diagram 2 Title: FtsZ Polymerization Cycle with Intervention Points
Conclusion
Reconstituting FtsZ and actin-like proteins within GUVs provides a powerful, reductionist platform to dissect the fundamental principles of bacterial cytoskeletal organization. By mastering the foundational concepts, methodological pipeline, and optimization strategies outlined here, researchers can generate robust, quantitative data that bridges in vitro biochemistry and cellular physiology. This approach not only advances our basic understanding of bacterial division and shape determination but also opens new avenues for high-throughput screening of antimicrobial compounds that specifically target these essential, dynamic protein assemblies. Future directions will involve increasing system complexity with multi-protein encapsulations and integrating these minimal systems with gene expression circuits to create truly functional synthetic cells.