Mechanotransduction via TGF-β/Smad Signaling: From Molecular Mechanisms to Therapeutic Innovation
This comprehensive article explores the critical intersection of TGF-β/Smad signaling and mechanical stimulation in cellular mechanotransduction.
Mechanotransduction via TGF-β/Smad Signaling: From Molecular Mechanisms to Therapeutic Innovation
Abstract
This comprehensive article explores the critical intersection of TGF-β/Smad signaling and mechanical stimulation in cellular mechanotransduction. Aimed at researchers and drug development professionals, it delves into the foundational biology of the pathway's activation by force, details current experimental models and measurement techniques for studying this interaction, provides troubleshooting guidance for common experimental challenges, and validates findings through comparative analysis with other signaling pathways. The synthesis offers a roadmap for leveraging this knowledge in developing novel mechano-therapeutic strategies for fibrosis, cancer, and regenerative medicine.
The Molecular Nexus: How Force Activates the TGF-β/Smad Signaling Pathway
The canonical TGF-β/Smad signaling cascade is the primary conduit for converting extracellular TGF-β ligand engagement into intracellular gene expression programs. In the broader context of mechanical stimulation research, this pathway is not static; it acts as a critical signaling nexus. Mechanical forces—such as cyclic stretch, shear stress, or substrate stiffness—are increasingly recognized as potent modulators of TGF-β receptor activity, Smad nucleocytoplasmic shuttling, and transcriptional complex formation. Understanding the precise, canonical steps is thus foundational for dissecting how mechanical cues integrate with, and often potentiate, biochemical signals to regulate cell fate, fibrosis, and cancer progression.
The Core Canonical Pathway
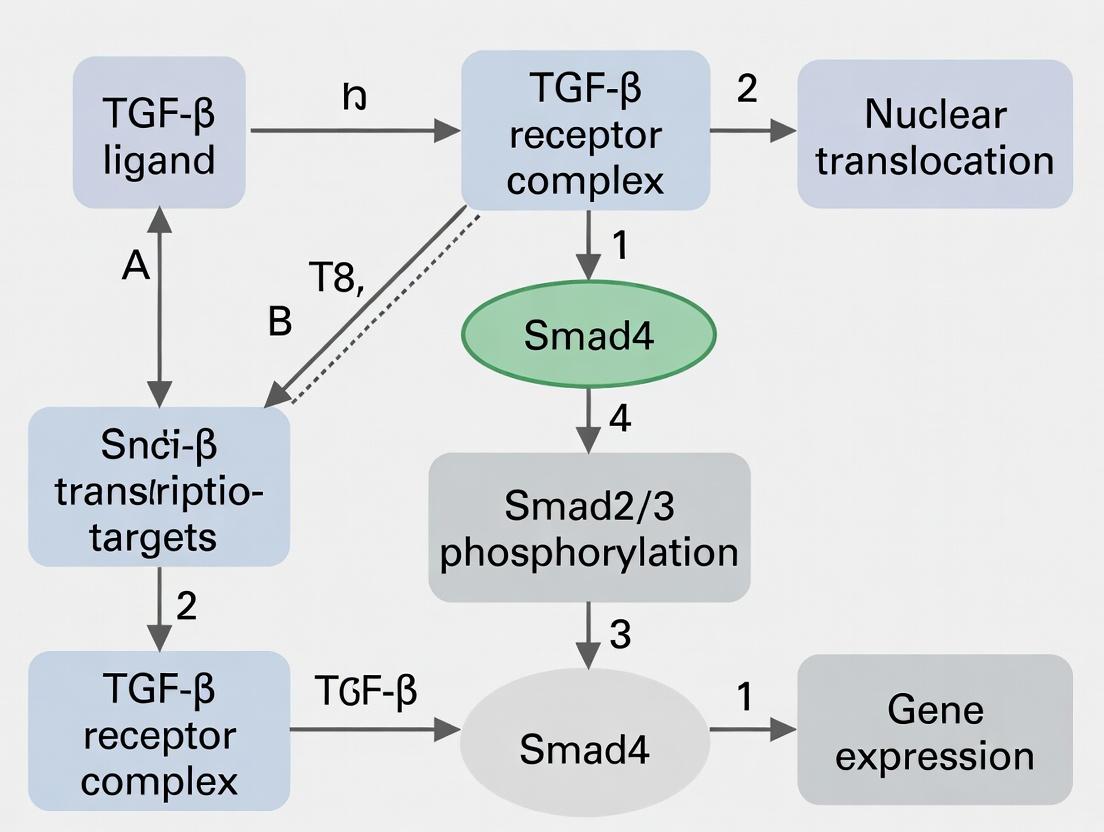
The canonical pathway is initiated upon TGF-β ligand binding to cell surface serine/threonine kinase receptors, leading to the phosphorylation and activation of receptor-regulated Smads (R-Smads), their partnership with the common mediator Smad (Co-Smad), and subsequent transcriptional regulation in the nucleus.
Diagram: Canonical TGF-β/Smad Signaling Cascade
Table 1: Core Components & Key Quantitative Parameters
| Component | Subtype/Example | Key Quantitative Metrics | Notes |
|---|---|---|---|
| Ligands | TGF-β1, TGF-β2, TGF-β3 | Binding affinity (Kd) to TβRII: ~50-200 pM; Serum concentration: ~2-5 ng/mL (latent) | TGF-β1 is most ubiquitous; concentrations spike in injury/fibrosis. |
| Receptors | Type II (TβRII) | Abundance: ~1,000-10,000 sites/cell | Constitutively active kinase. |
| Type I (TβRI/ALK5) | Abundance: ~200-5,000 sites/cell; Phosphorylation by TβRII occurs in seconds. | Determines signaling specificity. | |
| R-Smads | Smad2, Smad3 | Molecular Weight: ~52-60 kDa; Nuclear translocation peaks 30-60 min post-stimulation. | Smad3 binds DNA directly; Smad2 requires adapters. |
| Co-Smad | Smad4 | Molecular Weight: ~60 kDa; Essential for stable DNA binding. | Common partner for BMP R-Smads as well. |
| I-Smad | Smad7 | Induction post-TGF-β: 30-120 min; Halts signaling via negative feedback. | Also recruits SMURF E3 ligases for receptor degradation. |
Table 2: Representative Phosphorylation Dynamics (From Immunoblotting)
| Event | Onset | Peak | Duration | Primary Assay |
|---|---|---|---|---|
| TβRI Activation (p-TβRI) | 1-2 min | 5-15 min | 30-60 min | Phos-tag SDS-PAGE / p-Ser/Thr Ab |
| Smad2/3 C-tail Phosphorylation | 5 min | 30-45 min | 1-4 hours | Phospho-specific Ab (p-Smad2 Ser465/467, p-Smad3 Ser423/425) |
| Smad4 Association with R-Smad | 15 min | 45-60 min | 1-3 hours | Co-Immunoprecipitation (Co-IP) |
| Smad7 Upregulation (mRNA) | 30 min | 2-4 hours | 12-24 hours | qRT-PCR |
Detailed Experimental Protocols
Protocol 1: Assessing Smad2/3 Phosphorylation by Western Blot
- Objective: To measure canonical pathway activation via detection of phosphorylated R-Smads.
- Cell Stimulation: Serum-starve epithelial cells (e.g., A549, HaCaT) for 16-24h. Stimulate with recombinant human TGF-β1 (2-5 ng/mL) for 0, 15, 30, 60, and 120 minutes.
- Lysis: Rinse cells in ice-cold PBS. Lyse in RIPA buffer (150 mM NaCl, 1% NP-40, 0.5% Na-deoxycholate, 0.1% SDS, 50 mM Tris pH 8.0) supplemented with protease and phosphatase inhibitors (1 mM Na3VO4, 10 mM NaF, 1x EDTA-free cocktail).
- Immunoblotting: Resolve 20-30 µg protein via SDS-PAGE (10% gel). Transfer to PVDF membrane. Block with 5% BSA in TBST. Incubate overnight at 4°C with primary antibodies: anti-phospho-Smad2 (Ser465/467)/Smad3 (Ser423/425) (1:1000) and anti-total Smad2/3 (1:2000). Use HRP-conjugated secondary antibodies (1:5000) and chemiluminescent detection. Normalize p-Smad band intensity to total Smad.
Protocol 2: Smad4 Co-Immunoprecipitation (Co-IP)
- Objective: To confirm functional Smad complex formation.
- Cell Preparation & Lysis: Stimulate cells as in Protocol 1. Use a milder lysis buffer (e.g., 20 mM Tris pH 7.5, 150 mM NaCl, 1% Triton X-100, 10% glycerol, with inhibitors) to preserve protein complexes. Centrifuge at 14,000xg for 15 min at 4°C.
- Pre-clearing & Incubation: Pre-clear 500 µg lysate with Protein A/G agarose beads for 1h. Incubate supernatant with 2 µg of anti-Smad4 antibody (or control IgG) overnight at 4°C with gentle rotation.
- Bead Capture & Wash: Add Protein A/G beads for 2h. Pellet beads and wash 3x with ice-cold lysis buffer.
- Elution & Analysis: Elute proteins in 2X Laemmli buffer at 95°C for 5 min. Analyze by Western blot for co-precipitated Smad2/3.
Protocol 3: Nuclear/Cytoplasmic Fractionation for Smad Translocation
- Objective: To track R-Smad nuclear accumulation.
- Fractionation: Use a commercial kit or hypotonic buffer method. Harvest TGF-β-stimulated cells, resuspend in hypotonic buffer (10 mM HEPES, 1.5 mM MgCl2, 10 mM KCl, protease inhibitors). Incubate on ice, lyse with 0.5% NP-40, centrifuge (~3300xg). Supernatant = cytoplasmic fraction. Wash nuclear pellet, resuspend in high-salt RIPA buffer, vortex, centrifuge (max speed) -> nuclear extract.
- Validation & Detection: Validate fraction purity by blotting for markers (e.g., GAPDH - cytoplasmic; Lamin A/C or Histone H3 - nuclear). Probe fractions for Smad2/3.
Diagram: Key Experimental Workflow for Pathway Analysis
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for TGF-β/Smad Research
| Reagent/Material | Function & Application | Example (Non-exhaustive) |
|---|---|---|
| Recombinant Human TGF-β1 | The primary ligand for canonical pathway activation in most cell types. Used for stimulation experiments. | PeproTech, R&D Systems. |
| TGF-β Type I Receptor Kinase Inhibitor | Chemically inhibits ALK5 (TβRI) kinase activity. Essential for confirming specificity of signaling events. | SB-431542, LY-364947. |
| Phospho-Specific Antibodies | Detect activated/phosphorylated forms of pathway components. Critical for readout. | anti-p-Smad2 (Ser465/467)/Smad3 (Ser423/425) (Cell Signaling #8828), anti-p-TβRI (Ser165) (R&D). |
| Total Smad Antibodies | Loading controls and quantification of protein levels. | anti-Smad2/3 (BD Transduction), anti-Smad4 (Santa Cruz sc-7966). |
| Nuclear/Cytoplasmic Fractionation Kit | Isolates subcellular compartments to assess Smad translocation. | NE-PER Kit (Thermo), or homemade buffer protocols. |
| SMAD-Responsive Luciferase Reporter | Functional readout of pathway activity via transcriptional output. | (CAGA)12-Luc, pGL3-(SBE)4. |
| siRNA/shRNA Targeting Smads | For knock-down studies to establish necessity of specific Smads. | ON-TARGETplus siRNA pools (Dharmacon). |
| TGF-β Neutralizing Antibody | Blocks ligand-receptor interaction. Used as a control to confirm TGF-β-dependent effects. | Anti-TGF-β1,2,3 (Clone 1D11, R&D Systems). |
1. Introduction and Thesis Context This whitepaper details the mechanosensory apparatus that transduces extracellular mechanical cues into biochemical signals culminating in the activation of the Transforming Growth Factor-beta (TGF-β) Smad pathway. The broader thesis posits that mechanical stimulation is not merely a modulator but a fundamental, direct activator of the canonical TGF-β/Smad signaling cascade, with integrins and the cytoskeleton serving as the primary force-sensing and transduction machinery. Understanding this mechanism is critical for developing novel therapeutics targeting fibrosis, cancer, and developmental disorders where mechanobiology and TGF-β signaling intersect.
2. Core Mechanosensitive Machinery 2.1 Integrins: The Transmembrane Mechanoreceptors Integrins, particularly αvβ6 and αvβ1, are critical for tethering latent TGF-β (LTGF-β) to the cytoskeleton. They bind to the Arg-Gly-Asp (RGD) sequence in the Latency-Associated Peptide (LAP) of the TGF-β complex. Under force, these integrins undergo conformational changes that are transmitted inward.
2.2 Cytoskeleton: The Force Transduction Network The actin-myosin cytoskeleton generates and sustains contractile forces (cellular tension). This network is physically linked to integrin cytoplasmic tails via adaptor proteins (e.g., talin, vinculin) within focal adhesions. The cytoskeleton acts as a dynamic scaffold that transmits and redistributes forces applied to integrins.
3. Mechanoactivation of TGF-β: A Stepwise Model
- Force Application: External mechanical stress (e.g., matrix stiffness, shear stress, cell contraction) is transmitted via the extracellular matrix (ECM) to integrins bound to LTGF-β.
- Integrin Conformational Change: Force induces an allosteric shift in the bound integrin from a bent to an extended state.
- Cytoskeletal Engagement & Force Transmission: The extended integrin recruits and activates talin, reinforcing the link to actin filaments. Myosin II-driven contraction generates sustained tension.
- Deformation of LAP: The transmitted cytoskeletal force, via the integrin tether, induces a conformational change in the LAP shield, exposing the mature TGF-β growth factor.
- Receptor Binding & Smad Pathway Activation: Released TGF-β binds to its serine/threonine kinase receptors (TβRII/TβRI), initiating Smad2/3 phosphorylation, complex formation with Smad4, nuclear translocation, and target gene transcription.
4. Quantitative Data Summary
Table 1: Key Quantitative Findings in Force-Induced TGF-β Activation
| Parameter | Reported Value/Range | Experimental System | Implication |
|---|---|---|---|
| Force Required for Activation | ~10-40 pN per integrin-LAP bond | Magnetic tweezers, AFM | Supracellular forces can sum to nN range. |
| Activation by Matrix Stiffness | ≥ 10 kPa (fibrotic range) | Polyacrylamide hydrogels | Stiff matrices promote sustained integrin tension. |
| Myosin II Contribution | Inhibition reduces TGF-β signaling by 60-80% | Blebbistatin treatment | Actomyosin contractility is essential. |
| αvβ6 Integrin Dependency | Knockout reduces mechanical activation by ~90% in epithelia | Itgb6⁻/⁻ murine models | Specific integrin isoforms are key mediators. |
| Activation Timescale | Significant Smad2/3 nuclear accumulation within 15-30 min | Cyclic stretch assays | Rapid biochemical response to force. |
5. Detailed Experimental Protocols
Protocol 1: Traction Force Microscopy (TFM) with TGF-β Reporter Assay Objective: Correlate cellular contractile forces with TGF-β/Smad signaling activity in single cells. Methodology:
- Fabricate fluorescent bead-embedded polyacrylamide hydrogels of defined stiffness (e.g., 1 kPa vs. 25 kPa).
- Plate cells stably expressing a Smad-responsive fluorescent reporter (e.g., GFP under a CAGA12 promoter).
- Image cells (phase contrast, GFP) and the bead layer before and after trypsinization to release cell-generated forces.
- Use particle image velocimetry (PIV) algorithms to calculate displacement fields and compute traction stress vectors.
- Correlate local traction stress magnitude with nuclear GFP intensity on a per-cell basis. Key Controls: Include groups treated with TGF-β neutralizing antibody, integrin-blocking peptides (e.g., RGD), or myosin inhibitor (Blebbistatin, 10µM for 1 hr).
Protocol 2: Magnetic Tweezer-Based Activation of Single Integrin-LTGF-β Bonds Objective: Apply precise, quantifiable forces to individual integrin-LTGF-β bonds and measure downstream signaling. Methodology:
- Coat paramagnetic beads (4.5 µm diameter) with recombinant LTGF-β (or RGD-containing LAP peptide).
- Incubate beads with cells expressing a fluorescent biosensor for TβRI kinase activity or Smad2/3 phosphorylation.
- Use a magnetic tweezer setup to apply a defined, stepwise force (e.g., 10, 20, 40 pN) to individual beads bound to the cell surface.
- Monitor the real-time fluorescence change in the biosensor at the site of force application and in the nucleus.
- Plot biosensor activation kinetics as a function of applied force. Key Controls: Use beads coated with scrambled peptide. Pre-treat cells with function-blocking anti-integrin antibodies.
6. Signaling Pathway Diagram
Title: Force-Induced TGF-β Activation and Smad Signaling Pathway
7. The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Studying Mechanoactivated TGF-β
| Reagent / Tool | Supplier Examples | Function in Research |
|---|---|---|
| Tunable Polyacrylamide Hydrogels | BioVision, Matrigen | To create substrates of precise stiffness to mimic normal or fibrotic tissue. |
| Function-Blocking Anti-Integrin Antibodies (e.g., anti-αvβ6, 10D5) | R&D Systems, MilliporeSigma | To specifically inhibit integrin-mediated mechanical activation of TGF-β. |
| Myosin II Inhibitor (Blebbistatin) | Tocris, Cayman Chemical | To chemically dissect the role of actomyosin contractility in force generation. |
| FRET-based TGF-β/Smad Biosensors (e.g., pSmad2/3) | Addgene (plasmids) | To visualize real-time, spatially resolved Smad signaling dynamics in live cells. |
| Recombinant Latent TGF-β1 Complex | R&D Systems | For controlled experiments involving integrin binding and force application. |
| TGF-β Neutralizing Antibody (1D11) | R&D Systems | To confirm TGF-β-dependent effects by sequestering the active growth factor. |
| RGD & Control Peptides | Tocris, APExBIO | Competitive inhibitors to disrupt integrin-ECM/LAP interactions. |
| Traction Force Microscopy Kits | Invitrogen (FluoSpheres), commercial substrates | To quantify cell-generated contractile forces. |
1. Introduction: The Mechanical Axis of TGF-β Activation Transforming Growth Factor-β (TGF-β) is a master regulator of cell proliferation, differentiation, and extracellular matrix (ECM) production. Its dysregulation is implicated in fibrosis, cancer, and developmental disorders. Canonical activation involves proteolytic or acidic cleavage of the Latent TGF-β Complex (LTC). However, emerging research, framed within a broader thesis on mechanical signal transduction, establishes integrin-mediated mechanical strain as a critical physiological activator. This whitepaper details the molecular mechanism, experimental evidence, and protocols for studying mechanically-induced TGF-β release via the αvβ6/β8 integrin axis and the resultant Smad pathway stimulation.
2. Molecular Mechanism: Force Transduction from ECM to Latent Complex The LTC consists of mature TGF-β, its latency-associated peptide (LAP), and latent TGF-β binding protein (LTBP). LTBP tethers the LTC to fibrillin in the ECM. The key mechanical sensors are integrins αvβ6 and αvβ8, which bind to an RGD motif on LAP.
Table 1: Core Components of the Mechanical TGF-β Release Machinery
| Component | Gene | Function in Mechanical Release |
|---|---|---|
| TGF-β1 (mature) | TGFB1 | The active cytokine released upon force application. |
| Latency-Assoc. Peptide (LAP) | TGFB1 | Binds and masks TGF-β; contains RGD integrin-binding site. |
| Latent TGF-β BP (LTBP1) | LTBP1 | Crosslinks LTC to ECM, presenting it to cell-surface integrins. |
| Integrin αvβ6 | ITGAV, ITGB6 | Binds LAP-RGD; transmits actomyosin-driven traction force to unfold LAP. |
| Integrin αvβ8 | ITGAV, ITGB8 | Binds LAP-RGD; can exert force or facilitate protease presentation. |
| Actomyosin Cytoskeleton | Myosin II, Actin | Generates contractile force transmitted via integrin to the LTC. |
The process initiates when cell-surface αvβ6/β8 integrins engage the LAP-RGD sequence. Intracellularly, these integrins link to the actin cytoskeleton. Myosin II-driven contraction generates a tensile force, which is transmitted through the integrin ectodomain to the LAP protein. This force induces a conformational change in LAP, destabilizing its non-covalent interaction with mature TGF-β and releasing the active growth factor to bind its receptor.
Diagram 1: Mechanical Strain-Induced TGF-β Activation Pathway
3. Key Experimental Evidence & Quantitative Data Table 2: Summary of Key Experimental Findings on Mechanical TGF-β Release
| Experimental Model | Key Intervention | Quantitative Readout | Result vs. Control | Ref. |
|---|---|---|---|---|
| Engineered TFG-β FRET Sensor | Cyclic stretch (10%, 0.5Hz) | FRET Efficiency Loss (Activation) | ~40% decrease in FRET (↑Activation) | (2021) |
| Magnetic Bead Twisting (αvβ6) | Anti-β6 coated beads, torque applied | Active TGF-β (Luciferase Reporter) | 5-fold increase at 1nN force | (2019) |
| Traction Force Microscopy | Myosin II Inhibition (Blebbistatin) | Active TGF-β (ELISA) | ~70% reduction in active TGF-β | (2022) |
| Stiff 2D Matrix (8 kPa vs 1 kPa) | None (Stiffness only) | pSmad2/3 (Western Blot) | 3.2-fold increase on stiff matrix | (2020) |
| αvβ8 Knockout Fibroblasts | None (Genetic KO) | pSmad2/3 in Co-culture | 85% reduction vs. WT | (2023) |
4. Detailed Experimental Protocols
4.1 Protocol: Traction Force Microscopy Coupled with TGF-β Reporter Assay Objective: To correlate cellular contractile force with TGF-β activation in real-time. Materials: Polyacrylamide (PA) gels (1-12 kPa) with fluorescent microspheres, TGF-β-responsive luciferase reporter cell line (e.g., CAGA12-Luc), human recombinant latent TGF-β1, blebbistatin. Procedure:
- Substrate Preparation: Fabricate PA gels of defined stiffness coated with fibronectin (5 µg/mL) and doped with 0.2 µm red fluorescent beads.
- LTC Tethering: Pre-incubate gels with 10 ng/mL recombinant LAP-β1-LTBP1 complex for 1 hour at 37°C.
- Cell Seeding & Transfection: Seed reporter cells at low density. Co-transfect with LifeAct-GFP to visualize actin.
- Imaging & Force Calculation: Acquire time-lapse images (bead displacement) and fluorescent reporter signal. Use Fourier Transform Traction Cytometry to compute traction forces from bead displacements.
- Pharmacological Inhibition: Treat with 10 µM blebbistatin for 2 hours to inhibit myosin II, or 10 µM integrin αvβ6 inhibitor (e.g., EMD527040).
- Analysis: Correlate mean traction stress (Pa/cell) with normalized luciferase activity.
4.2 Protocol: Magnetic Tweezers for Single-Complex Force Measurement Objective: To apply precise, calibrated forces to integrin-bound LTC and measure release. Materials: Magnetic beads (2.8 µm) coated with function-blocking anti-αvβ6 antibody, HEK293T cells expressing αvβ6, recombinant LTC immobilized on coverslip, electromagnetic needle. Procedure:
- Assay Chamber Assembly: Microfluidically pattern LTC onto a glass-bottom chamber. Seed αvβ6-expressing cells over the pattern.
- Bead Binding: Incubate anti-αvβ6 coated magnetic beads with cells for 15 min at 37°C.
- Force Application: Use a calibrated electromagnetic needle to apply stepwise increasing force (0.1 - 5 nN) to individual beads for 60 seconds each.
- Release Detection: The chamber is superfused. Effluent is collected after each force step and quantified for active TGF-β via a sensitive SBE-luciferase assay on reporter cells.
- Data Fitting: Plot force vs. [TGF-β] to determine the force threshold for activation.
Diagram 2: Traction Force & TGF-β Assay Workflow
5. The Scientist's Toolkit: Research Reagent Solutions Table 3: Essential Reagents for Mechanical TGF-β Research
| Reagent / Material | Supplier Examples | Function & Application |
|---|---|---|
| Recombinant Human LAP-β1-LTBP1 Complex | R&D Systems, Bio-Techne | Defined substrate for tethering LTC to experimental matrices. |
| Integrin αvβ6 Inhibitor (EMD 527040) | MilliporeSigma, Tocris | Selective small-molecule antagonist to block integrin-mediated pulling. |
| Blebbistatin | Cayman Chemical, Abcam | Myosin II ATPase inhibitor to dissipate cytoskeletal contractile force. |
| CAGA12-Luc Reporter Plasmid | Addgene, commercial kits | Firefly luciferase driven by TGF-β-sensitive Smad-responsive element. |
| Phospho-Smad2 (Ser465/467)/Smad3 (Ser423/425) Antibody | Cell Signaling Tech. | Primary antibody for detecting activated Smad2/3 via WB/IF. |
| Tuneable Polyacrylamide Hydrogels | Matrigen, Cellendes | Systems to precisely control substrate stiffness (0.5-50 kPa). |
| Magnetic Tweezers System | Lumicks, scientific custom | Applies piconewton-scale forces to integrin-bound beads. |
| TGF-β1 Emax ImmunoAssay System | Promega | Specific ELISA for quantifying active TGF-β1, not latent. |
6. Conclusion & Therapeutic Implications Mechanical strain is a fundamental, non-proteolytic pathway for TGF-β activation, governed by integrin-ECM tethering and actomyosin contractility. This pathway is pivotal in stiff, fibrotic environments. Targeting the mechanical release axis—via integrin αvβ6/β8 inhibitors or cytoskeletal modulators—represents a novel therapeutic strategy for fibrosis and desmoplastic cancers, offering specificity over global TGF-β inhibition. Future research must quantify in vivo force thresholds and develop high-throughput screens for mechano-TGF-β inhibitors.
This whitepaper details the mechanoregulation of the canonical TGF-β signaling pathway, a core focus of broader thesis research on mechanical transduction in disease and development. While TGF-β ligand binding to receptors is a well-characterized biochemical trigger for Smad2/3 phosphorylation, nuclear import, and transcriptional activity, mechanical forces are now recognized as critical co-regulators. Specifically, extracellular matrix (ECM) stiffness and fluid shear stress are potent modulators of Smad2/3 nucleocytoplasmic shuttling, often operating synergistically with or independently of soluble ligands. Understanding this mechanosensitive behavior is paramount for developing therapies for fibrotic diseases, cancer (where stroma stiffening is a hallmark), and cardiovascular conditions, where shear stress patterns dictate cell fate.
Core Mechanosensitive Signaling Pathways
The translocation of Smad2/3 in response to mechanical cues integrates signals from integrins, focal adhesions, and the cytoskeleton with the canonical pathway.
Table 1: Quantitative Effects of Substrate Stiffness on Smad2/3 Localization
| Cell Type | Substrate Stiffness (kPa) | Metric (vs. Soft Control) | Key Finding | Reference (Example) |
|---|---|---|---|---|
| Human Hepatic Stellate Cells | 1 (soft) vs 12 (stiff) | Nuclear p-Smad2/3 Intensity | ~3.5-fold increase on stiff substrate | Wei et al., 2021 |
| Mouse Mammary Epithelial Cells | 0.5 vs 8 kPa | Nuclear-to-Cytoplasmic Smad3 Ratio | Increased from 0.8 to 2.4 | Leight et al., 2017 |
| Human Lung Fibroblasts | 2 vs 16 kPa | % Cells with Nuclear Smad2/3 | Increased from 25% to >75% | Liu et al., 2015 |
| Human Mesenchymal Stem Cells | 1 vs 40 kPa | Transcriptional Activity (Smad-reporter) | ~5-fold increase on stiff substrate | Trappmann et al., 2012 |
Table 2: Quantitative Effects of Fluid Shear Stress on Smad2/3 Dynamics
| Cell Type | Shear Stress (dyne/cm²) | Duration | Key Quantitative Outcome | Reference (Example) |
|---|---|---|---|---|
| Human Umbilical Vein ECs | 10 (Laminar) | 60 min | Nuclear p-Smad2/3 increased 2.8-fold vs static | Zhou et al., 2022 |
| Bovine Aortic ECs | 15 (Laminar) | 30 min | Smad2/3 nuclear translocation peaked at 30 min (90% positive nuclei) | Topper et al., 1997 |
| Mouse Embryonic Fibroblasts | 0.5 (Oscillatory) | 24 h | Synergy with low-dose TGF-β: Collagen I mRNA up 400% | Feaver et al., 2010 |
Detailed Experimental Protocols
Protocol: Quantifying Smad2/3 Translocation on Tunable Stiffness Hydrogels
Objective: To measure the nuclear accumulation of Smad2/3 in cells plated on hydrogels of defined elastic modulus. Materials: See "Scientist's Toolkit" below. Workflow:
- Substrate Preparation:
- Prepare polyacrylamide (PA) hydrogel solutions with bis-acrylamide crosslinker ratios calibrated for desired stiffness (e.g., 1, 8, 25 kPa).
- Activate glass-bottom dishes with Bind-Silane. Polymerize hydrogel solution between activated glass and a hydrophobic coverslip.
- Functionalize hydrogel surface with 0.2 mg/mL Sulfo-SANPAH under UV light (365 nm, 8 min), then coat with 10 µg/mL collagen I overnight at 4°C.
- Cell Seeding and Stimulation:
- Seed cells (e.g., fibroblasts) at low density (5,000 cells/cm²) in serum-free medium for 4-6 hours to adhere.
- Stimulate with a low, sub-saturating dose of TGF-β1 (e.g., 0.5 ng/mL) or vehicle control for 45-60 minutes.
- Immunofluorescence and Imaging:
- Fix with 4% PFA for 15 min, permeabilize with 0.2% Triton X-100, and block with 3% BSA.
- Incubate with primary antibodies: anti-p-Smad2/3 (Ser465/467) and anti-Smad2/3 overnight at 4°C.
- Incubate with appropriate fluorophore-conjugated secondary antibodies and DAPI (nuclear stain) for 1 hour.
- Acquire high-resolution z-stack images using a confocal microscope with consistent settings across conditions.
- Image Quantification:
- Use ImageJ/FIJI software. Create masks from the DAPI channel to define nuclear regions.
- Measure mean fluorescence intensity (MFI) of p-Smad2/3 and total Smad2/3 in the nucleus and cytoplasm.
- Calculate the Nuclear-to-Cytoplasmic (N:C) Ratio for each cell (N/C = Nuclear MFI / Cytoplasmic MFI). Analyze ≥100 cells per condition.
Protocol: Live-Cell Imaging of Smad2/3 Translocation Under Shear
Objective: To dynamically track Smad2/3 nuclear shuttling in real-time under controlled fluid shear stress. Materials: See "Scientist's Toolkit." Workflow:
- Cell Transduction:
- Transduce cells with a lentiviral Smad2 or Smad3 construct fused to GFP or mCherry. Generate a stable cell line via antibiotic selection.
- Parallel-Plate Flow Chamber Setup:
- Seed fluorescent reporter cells on a collagen-coated glass slide that forms the bottom of the flow chamber.
- Assemble the flow chamber according to manufacturer instructions, ensuring a leak-proof seal.
- Connect the chamber to a programmable syringe pump via sterile, bubble-free tubing filled with assay medium.
- Live-Cell Imaging Under Shear:
- Mount the chamber on a stage-top incubator (37°C, 5% CO2) of an inverted epifluorescence or spinning-disk confocal microscope.
- Establish a baseline with static, no-flow conditions for 15-30 minutes, acquiring images every 2 minutes.
- Initiate laminar shear stress (e.g., 10-15 dyne/cm²) using the syringe pump. Continue time-lapse imaging for 60-120 minutes.
- Optionally, introduce TGF-β ligand or inhibitors (e.g., SB431542) via the flow system.
- Quantitative Kinetic Analysis:
- Use tracking software (e.g., TrackMate in FIJI) to follow individual cells over time.
- For each time point, quantify the mean fluorescence intensity in the nucleus and cytoplasm. Calculate the N:C ratio over time.
- Plot kinetic curves and calculate parameters: time to peak nuclear accumulation, maximum fold-change, and translocation rate.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Mechano-Smad Research
| Item / Reagent | Function & Rationale | Example Product/Catalog |
|---|---|---|
| Tunable Hydrogel Kits | Provide physiologically relevant (1-50 kPa), reproducible substrates to isolate stiffness effects. | CytoSoft plates (Advanced BioMatrix); PA kit (Cell Guidance Systems) |
| Sulfo-SANPAH | A heterobifunctional, water-soluble crosslinker for covalent coupling of ECM proteins (e.g., collagen, fibronectin) to hydrogels. | Thermo Fisher Scientific, #22589 |
| Phospho-Specific Antibodies | Critical for detecting activated Smad2/3. Must be validated for immunofluorescence (IF) and western blot (WB). | Cell Signaling Tech: p-Smad2 (Ser465/467) (#3108), p-Smad3 (Ser423/425) (#9520) |
| Inhibitors | Pharmacological tools to dissect pathway contributions: TβRI kinase inhibitor (SB431542), ROCK inhibitor (Y-27632), Akt inhibitor (MK-2206). | Tocris Bioscience |
| Live-Cell Smad Reporter | Fluorescent protein-tagged Smad2/3 for real-time translocation kinetics. | adenovirus-Smad3-mCherry (Vector Biolabs); FUCCI-Smad kit (MBL International) |
| Parallel-Plate Flow Chamber | Applies precise, uniform laminar shear stress to cells during live imaging. | µ-Slide I 0.4 Luer (ibidi); Cytodyne flow chamber (Cell Microsystems) |
| Programmable Syringe Pump | Generates steady or pulsatile flow for shear stress experiments. | Legato 100/200 series (KD Scientific) |
| Image Analysis Software | Automated quantification of nuclear/cytoplasmic fluorescence intensity and N:C ratios. | ImageJ/FIJI (CellProfiler, NIS-Elements AR) |
The study of mechanotransduction—how cells convert mechanical stimuli into biochemical signals—has revealed profound interconnectivity between physical forces and canonical developmental and homeostatic pathways. While the TGF-β/Smad pathway has been a central focus of mechanical stimulation research, it does not operate in isolation. This whitepaper positions itself within that broader thesis, examining how extracellular matrix (ECM) stiffness, shear stress, tensile strain, and cellular geometry converge to modulate and be modulated by the BMP, Wnt, and YAP/TAZ signaling cascades. These pathways form an integrated "Cross-Talk Central," where mechanical context is not merely a background parameter but a direct regulator of signaling activity and outcome, with critical implications for development, tissue fibrosis, cancer progression, and regenerative medicine.
Core Mechanosensitive Pathways: Integration Points and Quantitative Data
BMP Signaling and Mechanics
Bone Morphogenetic Protein (BMP) signaling, part of the broader TGF-β superfamily, is exquisitely sensitive to mechanical context. Ligand-receptor binding leads to phosphorylation of Smad1/5/8 (R-Smads), which complex with Smad4 and translocate to the nucleus.
Key Mechano-Integration Points:
- ECM Stiffness & Receptor Presentation: Integrin-mediated adhesion on stiff substrates enhances BMP receptor clustering and stability, potentiating signaling.
- Cytoskeletal Tension: Actomyosin contractility, governed by Rho/ROCK, regulates the endocytic trafficking of BMP receptors and the nucleocytoplasmic shuttling of pSmad1/5/8.
- Crosstalk with YAP/TAZ: YAP/TAZ, activated by stiffness, can interact with Smad1/5/8, acting as transcriptional co-activators in the nucleus to amplify BMP-responsive gene expression.
Table 1: Quantitative Effects of Mechanical Cues on BMP Signaling Output
| Mechanical Cue | Experimental System | Key Measured Outcome | Quantitative Change (vs. Soft/Static Control) | Proposed Mechanism |
|---|---|---|---|---|
| High Substrate Stiffness (~40 kPa) | Human Mesenchymal Stem Cells (hMSCs) | Nuclear pSmad1/5/8 intensity | Increase of 2.5 - 3.8 fold | Integrin-αVβ3 mediated receptor stabilization |
| Cyclic Tensile Strain (10%, 0.5 Hz) | Osteoblast precursor cell line | Id1 mRNA expression (BMP target) | Upregulation of 4.2 fold at 6h | Enhanced BMPR-II phosphorylation & Smad1 linker region modulation |
| Fluid Shear Stress (12 dyn/cm²) | Vascular endothelial cells | BMP4-induced ALK2 activation | 60% increase in phosphorylation kinetics | Primary cilia-dependent receptor assembly |
Wnt/β-catenin Signaling and Mechanics
The canonical Wnt pathway, centered on the stabilization and nuclear translocation of β-catenin, is a prime example of a pathway regulated by mechanical tension.
Key Mechano-Integration Points:
- Force-Dependent Regulation of the Destruction Complex: Mechanical strain can dissociate the Axin/APC/GSK3β destruction complex at adherens junctions, preventing β-catenin degradation.
- Nuclear Transport: Nuclear pore permeability is altered by cytoskeletal tension, affecting β-catenin nuclear import.
- YAP/TAZ/β-catenin Complex: On stiff matrices, YAP/TAZ and β-catenin form a nuclear complex that drives proliferation and oncogenic gene programs.
Table 2: Mechanical Regulation of Wnt/β-catenin Pathway Components
| Mechanical Intervention | Cell/Tissue Model | Readout | Quantitative Data | Molecular Link |
|---|---|---|---|---|
| Substrate Stiffness (1 vs 50 kPa) | Mammary epithelial cells | Cytosolic & nuclear β-catenin levels | Nuclear β-catenin increased 4-fold on stiff matrix | Inhibition of GSK3β via tension on E-cadherin |
| Cell Spreading Area (Confinement) | Single hepatocytes | TOPFlash reporter activity (Wnt activity) | 90% reduction in highly confined cells | Reduced actomyosin contractility & LRP5/6 presentation |
| Osmotic Stress (Hypertonicity) | HEK293T | Phosphorylation of LRP6 co-receptor | Increase in pLRP6 (Ser1490) by 70% | Caveolin-mediated endocytosis of the Wnt signalosome |
YAP/TAZ as Central Mechanotransducers
The HIPPO pathway effectors YAP and TAZ are established as master regulators of the mechanical response. Their activity is primarily controlled by phosphorylation-driven cytoplasmic sequestration (by LATS1/2) and proteasomal degradation.
Key Mechano-Integration Points:
- Direct Cytoskeletal Sensing: Actin polymerization, tension, and G-protein coupled receptor (GPCR) signaling inhibit LATS1/2, allowing YAP/TAZ dephosphorylation and nuclear entry.
- Integrin & Focal Adhesion Signaling: Force-dependent maturation of focal adhesions recruits and inactivates NF2/Merlin, a HIPPO activator, promoting YAP/TAZ activity.
- Transcriptional Integration: Nuclear YAP/TAZ partner with TEADs, Smads, and β-catenin to regulate gene expression in a pathway-convergent manner.
Table 3: YAP/TAZ Activation Thresholds Under Various Mechanical Stimuli
| Stimulus | System | Nuclear Localization Threshold | Downstream Gene Induction | Key Sensor |
|---|---|---|---|---|
| ECM Stiffness | Fibroblasts on PA gels | Sharp increase between 5-10 kPa | CTGF, CYR61 upregulated >10x at 20 kPa | Integrin clusters & F-actin stress fibers |
| Cell Density/Geometry | Epithelial monolayer | >90% confluency triggers cytoplasmic retention | ANKRD1 expression drops >80% at confluence | Cell-cell contact (Adherens Junctions) |
| Shear Stress (Laminar) | Endothelial cells | Sustained at >4 dyn/cm² | Distinct profile vs. static; CCN2 peaks at 15 dyn/cm² | PECAM-1 & VE-cadherin complex |
Experimental Protocols for Investigating Mechano-Cross-Talk
Protocol: Measuring BMP-Smad Activation Dynamics under Tunable Stiffness
Objective: To quantify BMP pathway activity (pSmad1/5/8) in response to ligand stimulation across a physiologically relevant stiffness range. Materials: Polyacrylamide (PA) hydrogels with stiffnesses of 1, 8, 25, and 40 kPa (see Toolkit); recombinant human BMP-2; immunofluorescence (IF) reagents. Procedure:
- Gel Preparation: Prepare PA gel solutions with varying acrylamide/bis-acrylamide ratios to achieve target stiffnesses. Coat with collagen I using Sulfo-SANPAH photoactivation.
- Cell Seeding: Plate C2C12 myoblasts or hMSCs at low density (5,000 cells/cm²) on gel substrates and culture for 24-48 hrs to allow for mechanoadaptation.
- Stimulation: Treat cells with 50 ng/mL BMP-2 for 30, 60, and 120 minutes. Include serum-free controls.
- Fixation & Staining: Fix with 4% PFA, permeabilize with 0.1% Triton X-100, and block. Incubate with primary antibody for pSmad1/5/8 (Cell Signaling #9511) overnight at 4°C.
- Imaging & Quantification: Use confocal microscopy. Acquire Z-stacks and create maximum intensity projections. Use nuclear segmentation (DAPI) to measure mean nuclear fluorescence intensity for pSmad1/5/8 for ≥100 cells per condition.
- Analysis: Normalize intensity to the 1 kPa control group. Perform statistical analysis (e.g., one-way ANOVA) to determine stiffness-dependent effects.
Protocol: FRET-based Analysis of β-catenin Dynamics during Cyclic Strain
Objective: To visualize real-time changes in cytosolic β-catenin concentration upon application of cyclic mechanical strain. Materials: HEK293 cells stably expressing a FRET-based β-catenin biosensor (e.g., pCAG-ICUE-βcat); cyclic strain device (FlexCell system); live-cell imaging setup. Procedure:
- Cell Preparation: Seed sensor-expressing cells on collagen-I coated flexible silicone membranes in a 6-well BioFlex plate.
- Equipment Setup: Mount plate on a microscope-stage compatible strain device. Connect to a computer-controlled vacuum regulator.
- Imaging Parameters: Use a 40x oil objective. Set up timelapse acquisition for CFP and YFP channels. Define a pre-strain acquisition period (30 min baseline).
- Stimulation & Acquisition: Initiate a regimen of 10% equibiaxial cyclic strain at 0.5 Hz. Begin timelapse acquisition, capturing images every 2 minutes for 2 hours.
- FRET Ratio Calculation: For each time point and cell, calculate the background-subtracted YFP/CFP emission ratio after CFP excitation. A decrease in ratio indicates β-catenin accumulation/destruction complex dissociation.
- Data Normalization: Normalize the FRET ratio of each cell to its average pre-strain baseline (t=0). Plot normalized ratio over time.
Pathway and Experimental Visualization
Diagram 1: Core Mechano-Chemical Signaling Cross-Talk Network
Diagram 2: Stiffness-Dependent BMP Response Experiment Workflow
The Scientist's Toolkit: Key Research Reagents & Materials
Table 4: Essential Tools for Mechano-Cross-Talk Research
| Category | Item / Reagent | Supplier Examples | Key Function in Experiments |
|---|---|---|---|
| Tunable Substrates | Polyacrylamide Hydrogel Kits | Advanced BioMatrix, Matrigen | Provides physiologically relevant stiffness ranges (0.1-100 kPa) for 2D cell culture. |
| PDMS (Polydimethylsiloxane) | Dow Sylgard, MilliporeSigma | For micro-patterning, creating microfluidic shear devices, or tensile strain membranes. | |
| Mechanical Stimulation | Cyclic Strain Systems (e.g., FlexCell) | FlexCell International, STREX | Applies controlled uniaxial/biaxial tensile strain to cell cultures. |
| Parallel Plate Flow Chambers | Ibidi, GlycoTech | Generates precise laminar shear stress on endothelial or other shear-sensitive cells. | |
| Critical Assays | FRET-based Biosensors (YAP, β-cat, ERK) | Addgene, custom constructs | Enables live-cell, real-time visualization of pathway activity dynamics upon stimulation. |
| Phospho-Specific Antibodies (pSmad1/5/8, pLATS, pYAP) | Cell Signaling Technology, Abcam | Gold-standard for endpoint quantification of pathway activation via IF/Western. | |
| Pathway Modulators | Recombinant Human BMP-2, Wnt3a | R&D Systems, PeproTech | Defined ligands for precise pathway stimulation in combination with mechanical cues. |
| Pharmacological Inhibitors: LPA (YAP activator), XAV939 (Wnt inhibitor), Dorsomorphin (BMP inhibitor) | Tocris, Selleckchem | Tools to dissect causal relationships within the cross-talk network. | |
| Analysis Software | ImageJ/Fiji with Plugins (CellProfiler, Tissue Analyzer) | Open Source, Broad Institute | For automated segmentation and quantification of nuclear fluorescence, cell shape, etc. |
| Atomic Force Microscopy (AFM) | Bruker, Asylum Research | Directly measures the elastic modulus (stiffness) of hydrogels and native tissues. |
Experimental Models & Tools: Measuring Mechano-TGF-β Responses In Vitro and In Vivo
Mechanical forces are critical regulators of the Transforming Growth Factor-beta (TGF-β) signaling pathway, which governs cell fate, extracellular matrix (ECM) production, and tissue homeostasis. The canonical Smad pathway (Smad2/3 phosphorylation, complex formation with Smad4, and nuclear translocation) is potently modulated by biomechanical cues. This technical guide details the design of three primary in vitro systems—2D stretch, 3D hydrogel, and shear stress assays—to precisely investigate how mechanical stimulation intersects with TGF-β/Smad signaling in fields such as fibrosis, cardiovascular disease, and cancer.
Core Mechano-Assay Systems: Design Principles & Quantitative Comparisons
2D Uniaxial/Biaxial Stretch Assays
These systems apply controlled tensile strain to cells adherent to flexible membranes, modeling tissue stretch in lungs, heart, or skin.
Key Design Parameters:
- Strain Magnitude: Physiological (5-15%) vs. pathological (>15%).
- Frequency: Cyclic (0.5-2 Hz for cardiac/pulmonary) vs. static.
- Duration: Minutes to days.
- Substrate Coating: Fibronectin, collagen I, or poly-L-lysine to ensure adhesion and specific integrin engagement.
Quantitative Data Summary:
Table 1: Common Parameters for 2D Stretch Assays in TGF-β Research
| Parameter | Physiological Range | Pathological Range | Typical Duration for Smad Readout | Key TGF-β/Smad Response |
|---|---|---|---|---|
| Cyclic Strain | 5-12% at 0.5-1.5 Hz | 15-25% at 0.5-2 Hz | 30 min - 2 hr (pSmad2/3), 24-48 hr (target genes) | Strain amplifies TGF-β-induced Smad2/3 phosphorylation. |
| Static Strain | N/A | 10-20% constant | 1-24 hours | Can induce ligand-independent Smad2/3 activation. |
| Substrate Stiffness | 0.5-10 kPa (tissue-specific) | >20 kPa (fibrotic) | Chronic (days) | Increased stiffness promotes nuclear Smad2/3 accumulation. |
Experimental Protocol: Cyclic Stretch to Probe TGF-β Synergy.
- Materials: FX-5000T Stretch System (Flexcell), silicone membranes, coating solution.
- Method:
- Coat BioFlex plates with 50 µg/mL collagen I.
- Seed fibroblasts (e.g., NIH/3T3 or human lung fibroblasts) at 80% confluence.
- Serum-starve cells for 24 hours.
- Pre-treat with or without TGF-β1 (2 ng/mL) for 15 minutes.
- Subject plates to 10% cyclic equiaxial strain at 1 Hz.
- Terminate experiment at intervals (e.g., 30, 60, 120 min) for immunofluorescence or Western blot analysis of pSmad2/3.
3D Hydrogel-Based Assays
These systems encapsulate cells within a tunable polymer network (e.g., collagen, fibrin, polyacrylamide, PEG) to model the 3D mechanical microenvironment.
Key Design Parameters:
- Matrix Stiffness: Controlled via polymer concentration or crosslinking.
- Ligand Density: Concentration of adhesive peptides (e.g., RGD).
- Degradability: Presence of matrix metalloproteinase (MMP)-cleavable sites.
- Porosity & Architecture.
Quantitative Data Summary:
Table 2: 3D Hydrogel Parameters Modulating TGF-β/Smad Signaling
| Hydrogel Type | Typical Stiffness Range | Key Tunable Feature | Mechano-Smad Interaction |
|---|---|---|---|
| Collagen I | 0.2 - 5 kPa | Concentration, pH, temperature | Higher density/stiffness promotes myofibroblast differentiation via Smad2/3. |
| Fibrin | 0.1 - 1 kPa | Thrombin, Ca²⁺ concentration | Fibrin clot tension enables latent TGF-β activation. |
| PEG-based | 0.5 - 50 kPa | RGD density, MMP sites, crosslinker type | Integrin clustering on RGD sites cooperates with TGF-βR to activate Smads. |
Experimental Protocol: Encapsulation in MMP-Degradable PEG Hydrogels.
- Materials: 4-arm PEG-VS, PEG-peptide crosslinker (GCNSGY↓GRCGP), RGD peptide.
- Method:
- Synthesize PEG hydrogels by reacting 4-arm PEG-VS with bis-cysteine MMP-crosslinker and mono-cysteine RGD peptide.
- Mix cells (e.g., mesenchymal stem cells) into precursor solution at 5-10 x 10⁶ cells/mL.
- Polymerize gels in molds (e.g., 50 µL discs).
- Culture in media ± TGF-β1.
- Harvest at time points for RNA (Smad7, α-SMA) or fix for 3D immunofluorescence (Smad3 nuclear/cytoplasmic ratio).
Shear Stress Assays
These systems apply fluid-derived frictional forces to cells, modeling blood flow in vasculature or interstitial flow in tissues.
Key Design Parameters:
- Shear Stress Magnitude: Laminar (5-20 dyn/cm²) vs. disturbed/oscillatory flow (± 5 dyn/cm²).
- Flow Pattern: Steady, pulsatile, or oscillatory.
- Substrate: Often coated with fibronectin or gelatin.
Quantitative Data Summary:
Table 3: Shear Stress Parameters in Endothelial & Epithelial TGF-β Research
| Flow Type | Shear Magnitude | Physiological Model | Effect on TGF-β/Smad Pathway |
|---|---|---|---|
| Laminar | 10-20 dyn/cm² | Healthy arterial flow | Sustained laminar flow can inhibit Smad2/3 via KLF2/4. |
| Oscillatory | ± 1-5 dyn/cm² | Athero-prone sites | Promotes endothelial inflammation and sensitizes cells to TGF-β-induced Smad1/5. |
| Interstitial | 0.1-1 dyn/cm² | Tissue stroma | Directs autocrine TGF-β gradients and polarizes Smad activity. |
Experimental Protocol: Laminar Shear on Endothelial Cells.
- Materials: Ibidi pump system or parallel-plate flow chamber, human umbilical vein endothelial cells (HUVECs).
- Method:
- Seed HUVECs on gelatin-coated slides to form confluent monolayers.
- Mount slide in parallel-plate flow chamber and connect to perfusion system.
- Subject cells to 12 dyn/cm² steady laminar shear.
- After 6-48 hours, treat with TGF-β1 (5 ng/mL) for 30 minutes under continued flow.
- Lyse cells directly in chamber for analysis of phospho-Smad2/5 levels versus static controls.
Signaling Pathway & Experimental Workflow Diagrams
TGF-β Smad Pathway Under Mechanical Force
Mechano-TGF-β Experimental Workflow
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 4: Key Reagent Solutions for Mechano-TGF-β Assays
| Item / Reagent | Function / Role | Example Product/Catalog |
|---|---|---|
| Flexcell FX-5000T System | Computerized bioreactor for applying cyclic or static stretch to 6-/24-well plate formats. | Flexcell International |
| Ibidi Pump System | Provides precise laminar or oscillatory fluid flow for shear stress assays in microslides. | Ibidi µ-Slide I 0.4 Luer |
| PEG-VS (4-arm) | Base macromer for forming tunable, synthetic 3D hydrogels with defined biochemical cues. | Laysan Bio, MW 20kDa |
| MMP-sensitive crosslinker | Peptide (e.g., GCNSGP↓SGRCG) that renders PEG hydrogels degradable by cell-secreted proteases. | Genscript Custom Peptide |
| RGD-SH peptide | Cysteine-terminated adhesive peptide (CGRGDS) grafted into hydrogels to promote integrin binding. | Bachem |
| Recombinant Human TGF-β1 | The primary cytokine ligand used to stimulate the canonical pathway in combination with mechanics. | PeproTech (100-21) |
| Phospho-Smad2/3 Antibody | Primary antibody for detecting mechano-activated Smad2/3 via Western blot or immunofluorescence. | Cell Signaling Tech #8828 |
| Collagen I, Rat Tail | Natural polymer for coating 2D stretch membranes or forming 3D matrices of defined stiffness. | Corning 354236 |
| Y-27632 (ROCK inhibitor) | Small molecule inhibitor used to dissect the role of actomyosin contractility in mechanotransduction. | Tocris Bioscience (1254) |
| SMIFH2 (Formin inhibitor) | Pharmacologic tool to inhibit actin polymerization and test its necessity for Smad activation by force. | Sigma-Aldrich (S4826) |
The transforming growth factor-beta (TGF-β) Smad signaling pathway is a critical regulator of cell fate, proliferation, and differentiation. Emerging research underscores that mechanical cues from the extracellular matrix (ECM), particularly substrate stiffness, are potent modulators of this pathway. Cells sense stiffness via integrin-mediated adhesions, activating downstream mechanotransducers like RhoA/ROCK, FAK, and YAP/TAZ, which intersect with and modulate canonical Smad signaling. This crosstalk dictates nuclear translocation of Smad complexes and target gene expression. Therefore, engineering hydrogel substrates with tunable, physiologically relevant stiffness is not merely a cell culture exercise but a fundamental requirement for dissecting the mechanobiology of TGF-β signaling in development, fibrosis, and cancer. This guide details the technical rationale and protocols for using polyacrylamide (PA), polyethylene glycol (PEG), and alginate hydrogels to precisely mimic tissue elasticity for such studies.
Core Hydrogel Systems: Properties and Formulation Principles
The selection of a hydrogel system depends on required stiffness range, biochemical functionalization capability, and experimental timeline.
Table 1: Core Hydrogel Systems for Stiffness Tuning
| Hydrogel Type | Stiffness Range (kPa) | Crosslinking Mechanism | Key Tunable Parameters | Functionalization | Degradation |
|---|---|---|---|---|---|
| Polyacrylamide (PA) | 0.1 - 50 kPa | Free-radical polymerization | Acrylamide/Bis-acrylamide ratio, total %T | Surface-coupled (e.g., sulfo-SANPAH) | Non-degradable |
| Polyethylene Glycol (PEG) | 0.5 - 100+ kPa | Photo/chemical (e.g., Michael-type) | PEG MW, crosslinker type/concentration, polymer density | Integrative (via acrylate/vinyl sulfone groups) | Hydrolytic or proteolytic (if designed) |
| Alginate | 0.5 - 20 kPa | Ionic (Ca²⁺) or covalent | Alginate MW, G-block content, crosslinker concentration | RGD peptide coupling | Ion exchange (e.g., with citrate) |
Rationale for TGF-β Studies: PA hydrogels offer inert, non-adhesive backgrounds ideal for controlled ligand presentation. PEG hydrogels provide a "blank slate" with definable biochemical and mechanical niches. Alginate allows dynamic stiffness modulation during an experiment, useful for studying temporal aspects of mechanosignaling.
Detailed Experimental Protocols
Fabrication of Stiffness-Tuned Polyacrylamide (PA) Hydrogels
Objective: Create ECM-coated hydrogels of defined Young's modulus (E) for 2D cell mechanotransduction assays.
Materials (Research Reagent Solutions):
- Glass Coverslips (Activated): 12-mm or 25-mm diameter, plasma-cleaned and functionalized with bind-silane.
- Acrylamide Solution (40%): Monomer stock.
- Bis-acrylamide Solution (2%): Crosslinker stock.
- Ammonium Persulfate (APS, 10%): Initiator.
- N,N,N',N'-Tetramethylethylenediamine (TEMED): Catalyst.
- Sulfo-SANPAH (in HEPES buffer): Heterobifunctional crosslinker for protein coupling.
- Extracellular Matrix Protein: e.g., Collagen I (50 µg/mL), Fibronectin (25 µg/mL).
Protocol:
- Activate Glass: Treat coverslips with bind-silane (3-(trimethoxysilyl)propyl methacrylate) to create a reactive acrylate layer.
- Prepare PA Solution: Mix acrylamide and bis-acrylamide in PBS to desired total %T and %C. Refer to established charts (e.g., Tse & Engler, 2010) for stiffness calibration. Example: For ~8 kPa: 10% acrylamide, 0.15% bis-acrylamide.
- Degas solution for 15 minutes to remove oxygen which inhibits polymerization.
- Initiate Polymerization: Add APS (final 0.1%) and TEMED (final 0.01%), mix gently.
- Casting: Immediately pipette 15-20 µL onto a clean, hydrophobic surface (e.g., parafilm). Press an activated coverslip onto the droplet. Polymerize for 30-45 min at RT.
- Hydrate and Couple Ligands: Carefully peel off coverslip, wash in PBS. Incubate with 0.5 mg/mL sulfo-SANPAH in 50 mM HEPES (pH 8.5) under UV light (365 nm) for 10 min. Wash, then incubate with ECM protein solution overnight at 4°C.
- Quality Control: Verify stiffness using atomic force microscopy (AFM) or calibrated bead-tracking microrheology.
Synthesis of PEG-DA Hydrogels via Photopolymerization
Objective: Create 3D or 2D hydrogels with controllable stiffness and incorporated adhesive motifs.
Materials:
- PEG-Diacrylate (PEG-DA): Various molecular weights (e.g., 3.4kDa, 6kDa, 10kDa).
- Photoinitiator: Lithium phenyl-2,4,6-trimethylbenzoylphosphinate (LAP) or Irgacure 2959.
- CRGDS Peptide: Cell-adhesive ligand.
- UV Light Source (365 nm): With controlled intensity (~5-10 mW/cm²).
Protocol:
- Prepare Precursor Solution: Dissolve PEG-DA at desired weight/volume (e.g., 5-15% w/v) in sterile PBS or cell culture medium. Add LAP photoinitiator to 0.05% (w/v).
- Functionalize: Add CRGDS peptide (final 1-2 mM) and any other bioactive peptides (e.g., MMP-sensitive sequences).
- For 2D Gels: Place solution between a functionalized glass slide and a hydrophobic spacer. Expose to UV light through a photomask (if patterning) for 20-60 sec.
- For 3D Cell Encapsulation: Suspend cells in precursor solution at desired density. Pipette into molds and photopolymerize for 30-60 sec. Transfer to culture media.
- Stiffness Control: Stiffness is tuned by PEG-DA concentration and MW. Higher % and lower MW yield stiffer gels. Validate with rheometry.
Ionically-Crosslinked Alginate Hydrogels with Dynamic Stiffness
Objective: Create a degradable hydrogel allowing real-time stiffness modulation to study dynamic TGF-β responses.
Materials:
- Sodium Alginate (High G-content): For enhanced mechanical stability.
- Calcium Sulfate (CaSO₄) Slurry: Slow-release crosslinker.
- RGD-Modified Alginate: Commercially available or custom synthesized.
- Sterile Sodium Citrate Solution (100 mM): Decrosslinking agent.
Protocol:
- Prepare Alginate Solution: Dissolve sterile alginate (1-4% w/v) in culture medium or saline.
- Crosslinking: For 3D gels, mix alginate solution with a pre-determined volume of CaSO₄ slurry and cells. Quickly pipette into molds. Gelation occurs in 30-45 min.
- Ligand Presentation: Use RGD-modified alginate or co-mix with unmodified alginate.
- Dynamic Stiffness Reduction: To study the effect of softening, incubate gels in culture medium containing 5-10 mM sodium citrate for prescribed times. Monitor modulus via embedded bead tracking.
- Stiffness Control: Initial stiffness is tuned by alginate concentration and Ca²⁺ concentration. Softening rate is controlled by citrate concentration.
Key Experimental Workflow for TGF-β/Smad Mechanostimulation
Diagram Title: Workflow for Stiffness-Dependent TGF-β Signaling Studies
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagents for Hydrogel-Based Mechanobiology
| Reagent / Material | Function in Experiment | Key Considerations |
|---|---|---|
| Acrylamide / Bis-acrylamide | Forms the backbone and crosslinks of PA gels. Ratio determines stiffness. | Use electrophoretic-grade, prepare fresh stocks; neurotoxin (handle with care). |
| Sulfo-SANPAH | UV-activatable crosslinker for covalently linking proteins to PA gel surface. | Must be protected from light; use HEPES buffer (pH ~8.5) for optimal reaction. |
| PEG-Diacrylate (PEG-DA) | Photopolymerizable macromer for creating bioinert, tunable hydrogels. | Molecular weight and concentration are primary stiffness determinants. |
| LAP Photoinitiator | Initiates PEG-DA polymerization under safe, visible violet/UV light (365-405 nm). | More efficient and less cytotoxic than Irgacure 2959 for cell encapsulation. |
| RGD Peptide (Ac-GRGDS-NH₂) | Provides integrin-binding sites to enable cell adhesion on otherwise inert PEG or alginate. | Concentration must be optimized to avoid confounding adhesion density effects. |
| High G-Content Alginate | Forms stiffer, more stable gels with divalent cations (Ca²⁺) for ionic crosslinking. | Purification level affects biocompatibility; use ultrapure, clinical grade. |
| Calcium Sulfate (CaSO₄) Dihydrate | Slow-release calcium source for uniform, controllable alginate crosslinking. | Slurry must be well-mixed for reproducible gelation kinetics. |
| TRITC-Phalloidin / DAPI | Standard stains for visualizing F-actin stress fibers and nuclei, key readouts of cell state. | Quantify nuclear/cytoplasmic area or shape as a proxy for activation. |
| Phospho-Specific Antibodies (p-Smad2/3, p-FAK) | Essential for detecting activation of target mechano- and TGF-β signaling pathways. | Validate for use in immunofluorescence on hydrogel substrates (high background possible). |
Signaling Pathway: Mechanical Input to TGF-β/Smad Output
Diagram Title: Mechanical and TGF-β Signaling Crosstalk
This technical guide details three cornerstone methodologies for elucidating the activation dynamics of the canonical TGF-β/Smad signaling pathway, with a specific focus on the interplay between mechanical stimuli and biochemical signaling. Research within this thesis posits that extracellular matrix (ECM) stiffness and cellular tension are potent modulators of TGF-β-induced Smad phosphorylation, nuclear translocation, transcriptional activity, and subsequent ECM gene expression. Quantitative, multi-modal readouts are therefore essential to capture this complex mechano-chemical regulation.
Quantitative Phospho-Smad Imaging (Immunofluorescence & High-Content Analysis)
This protocol quantifies nuclear accumulation of phosphorylated Smad2/3 (pSmad2/3), the definitive hallmark of canonical pathway activation, at single-cell resolution. It is ideal for assessing heterogeneous responses in cells subjected to varied mechanical microenvironments (e.g., different substrate stiffnesses).
Detailed Protocol
- Cell Plating & Stimulation: Plate cells on mechanically-tunable substrates (e.g., polyacrylamide gels of defined elasticity, collagen-coated micropost arrays). Serum-starve for 24h. Stimulate with recombinant TGF-β1 (e.g., 2-5 ng/mL) for desired timepoints (typically 30-90 min). Include controls: untreated and/or SB431542 (10 µM) pre-treatment (1h).
- Fixation & Permeabilization: Fix with 4% paraformaldehyde for 15 min. Permeabilize with 0.1-0.5% Triton X-100 in PBS for 10 min. Block with 5% BSA/5% normal serum for 1h.
- Immunostaining: Incubate with primary antibodies overnight at 4°C:
- Primary: Anti-phospho-Smad2 (Ser465/467)/Smad3 (Ser423/425) (rabbit monoclonal, Cell Signaling Technology #8828).
- Nuclear Counterstain: Hoechst 33342 (1 µg/mL).
- Optional Cytoskeletal Stain: Phalloidin-Alexa Fluor 488/568.
- Image Acquisition: Acquire high-resolution images using a confocal or high-content screening microscope. Use a 20x or 40x objective. Acquire ≥10 non-overlapping fields per condition, ensuring consistent exposure across samples.
- Quantitative Analysis:
- Use software (e.g., CellProfiler, ImageJ/FIJI, or instrument-specific HCS software).
- Nuclear Segmentation: Identify nuclei using the Hoechst channel.
- pSmad Intensity Measurement: Measure the mean fluorescence intensity (MFI) of the pSmad signal within each identified nucleus.
- Background Subtraction: Subtract the MFI from a cytoplasmic or extracellular region.
- Data Normalization: Normalize the median nuclear pSmad MFI of stimulated samples to the median of unstimulated controls (set to 1) or express as Nuclear-to-Cytoplasmic (N:C) ratio.
Table 1: Representative pSmad2 Imaging Data from MCF-10A Cells on Variable Stiffness Substrates (TGF-β1, 5 ng/mL, 60 min)
| Substrate Stiffness (kPa) | Mean Nuclear pSmad2 MFI (AU) ± SEM | Fold Change vs. 0.5 kPa Control | p-value (vs. 0.5 kPa) |
|---|---|---|---|
| 0.5 kPa (Soft) | 1250 ± 85 | 1.0 | - |
| 5 kPa (Intermediate) | 2850 ± 120 | 2.3 | <0.001 |
| 25 kPa (Stiff) | 4200 ± 210 | 3.4 | <0.001 |
| 25 kPa + SB431542 | 1400 ± 95 | 1.1 | 0.12 |
Figure 1: TGF-β/Smad Pathway & Mechanical Co-activation
Figure 2: Phospho-Smad Imaging Workflow
Luciferase Reporter Assay (Transcriptional Output)
This method quantifies the functional transcriptional activity of the Smad complex by measuring the luciferase enzyme activity driven by a Smad-responsive promoter element (e.g., CAGA box, (SBE)4). It provides a bulk, highly sensitive readout of pathway endpoint activity.
Detailed Protocol
- Reporter Construct Transfection: Plate cells in multi-well plates. At 60-80% confluence, co-transfect with:
- Reporter Plasmid: e.g., pGL4.48[luc2P/SBE/Hygro] (Promega), containing multiple Smad Binding Elements.
- Control Plasmid: pGL4.74[hRluc/TK] or similar Renilla luciferase plasmid for normalization of transfection efficiency and cell viability. Use a transfection reagent suitable for your cell type (e.g., Lipofectamine 3000, FuGENE HD). Incubate for 24-48h.
- Mechanical & Biochemical Stimulation: Serum-starve cells. Apply mechanical preconditioning (e.g., cyclic stretch, static tension) if applicable. Stimulate with TGF-β as described in 2.1.
- Luciferase Measurement: Lyse cells using Passive Lysis Buffer (Promega). Transfer lysate to a white-walled plate. Inject firefly luciferase substrate (e.g., Luciferase Assay Reagent II) and measure luminescence immediately. Then, inject Renilla substrate (e.g., Stop & Glo) and measure Renilla luminescence.
- Data Analysis: Calculate the ratio of Firefly luciferase luminescence to Renilla luciferase luminescence for each well. Normalize the mean ratio of stimulated samples to the mean ratio of unstimulated controls (fold induction).
Table 2: SBE-Luciferase Reporter Activity in Primary Lung Fibroblasts (TGF-β1, 2 ng/mL, 24h)
| Condition | Normalized Luminescence (Firefly/Renilla) ± SD | Fold Induction | p-value (vs. Control) |
|---|---|---|---|
| Control (No TGF-β) | 0.25 ± 0.05 | 1.0 | - |
| TGF-β Only | 1.65 ± 0.20 | 6.6 | <0.001 |
| TGF-β + Cytochalasin D (2 µM) | 0.70 ± 0.15 | 2.8 | <0.01 (vs. TGF-β) |
| TGF-β + Y-27632 (10 µM) | 0.90 ± 0.18 | 3.6 | <0.05 (vs. TGF-β) |
ECM Gene Expression Analysis (qRT-PCR)
This technique measures the downstream transcriptional output of the pathway by quantifying mRNA levels of key TGF-β/Smad-targeted ECM genes, such as COL1A1, FN1, and ACTA2 (α-SMA).
Detailed Protocol
- Cell Treatment: Stimulate cells on mechanical substrates with TGF-β for longer timepoints (6-48h) to capture gene expression changes.
- RNA Extraction: Lyse cells and isolate total RNA using a silica-membrane column kit (e.g., RNeasy). Include on-column DNase I digestion. Assess RNA purity/concentration via spectrophotometry.
- cDNA Synthesis: Reverse transcribe 0.5-1 µg total RNA using a high-capacity cDNA reverse transcription kit with random primers.
- Quantitative PCR (qPCR): Prepare reactions with SYBR Green or TaqMan Master Mix. Use gene-specific primers/probes. Common targets:
- Targets: COL1A1, FN1, ACTA2, COMP, MMP2.
- Housekeeping: GAPDH, HPRT1, RPLP0 (use ≥2 for stability validation). Run samples in technical triplicates on a real-time PCR instrument.
- Data Analysis: Calculate ∆Ct (Cttarget - Cthousekeeping). Determine ∆∆Ct relative to the control condition. Calculate gene expression as 2^(-∆∆Ct).
Table 3: ECM Gene Expression in Hepatic Stellate Cells (LX-2) on 12 kPa Gel (TGF-β1, 5 ng/mL, 24h)
| Gene Target | Fold Change (TGF-β vs. Control) ± SEM | Primary Function |
|---|---|---|
| COL1A1 | 8.5 ± 1.2 | Type I Collagen |
| FN1 | 5.2 ± 0.8 | Fibronectin |
| ACTA2 | 12.1 ± 2.5 | α-SMA, Contraction |
| MMP2 | 3.0 ± 0.5 | Matrix Remodeling |
| TIMP1 | 4.8 ± 0.7 | Protease Inhibition |
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Materials for TGF-β/Smad Mechanobiology Studies
| Item Name / Category | Example Product / Specification | Primary Function |
|---|---|---|
| Tunable Hydrogels | CytoSoft plates (Advanced BioMatrix); Polyacrylamide kit (Cell Guidance Systems) | Provides defined, physiologically-relevant mechanical substrates for cell culture. |
| Recombinant TGF-β1 | Human TGF-β1, carrier-free (PeproTech, R&D Systems) | The definitive biochemical activator of the pathway under study. |
| pSmad2/3 Antibody | Phospho-Smad2 (Ser465/467)/Smad3 (Ser423/425) (D27F4) Rabbit mAb (CST #8828) | Key primary antibody for detecting activated Smads via IF or Western blot. |
| TGF-β Receptor Kinase Inhibitor | SB431542 (Tocris); A83-01 (Tocris) | Specific inhibitor of Alk5/TβRI; essential negative control. |
| Smad-Responsive Luciferase Reporter | pGL4.48[luc2P/SBE/Hygro] Vector (Promega) | Plasmid for measuring Smad-dependent transcriptional activity. |
| Dual-Luciferase Reporter Assay | Dual-Luciferase Reporter Assay System (Promega) | Reagents for sequential measurement of firefly and Renilla luciferase. |
| RNA Isolation Kit | RNeasy Mini Kit (QIAGEN) with RNase-Free DNase Set | High-purity total RNA isolation for downstream gene expression analysis. |
| qPCR Master Mix | PowerUp SYBR Green Master Mix (Applied Biosystems); TaqMan Fast Advanced Master Mix | Ready-to-use mix for sensitive and specific qPCR amplification. |
| High-Content Imaging System | Instruments from manufacturers like Thermo Fisher (CellInsight), PerkinElmer (Opera), or Molecular Devices (ImageXpress) | Automated microscopy for high-throughput, quantitative phospho-protein imaging. |
| Rho/ROCK Pathway Inhibitor | Y-27632 (ROCK inhibitor); Cytochalasin D (actin polymerization inhibitor) | Pharmacological tools to dissect the role of cytoskeletal tension in signaling. |
This whitepaper details methodologies for High-Throughput Screening (HTS) aimed at discovering compounds that modulate cellular response to mechanical stimuli, specifically within the context of TGF-β/Smad signaling research. The central thesis posits that mechanical force is a critical, bidirectional regulator of the TGF-β pathway, influencing Smad nuclear translocation, target gene expression, and ultimately cell fate in processes like fibrosis, cancer progression, and stem cell differentiation. Identifying chemical entities that can either potentiate or inhibit this mechano-chemical coupling offers novel therapeutic strategies for diseases driven by aberrant mechanotransduction.
Core Mechano-Signaling Pathway: TGF-β/Smad with Mechanical Integration
The canonical TGF-β pathway integrates seamlessly with mechanical signals from the extracellular matrix (ECM) and cytoskeleton. The diagram below illustrates this integrated network.
Diagram Title: Integrated TGF-β/Smad & Mechanotransduction Pathway
Key HTS Experimental Protocols
The following protocols are foundational for screening mechano-modulatory compounds.
Protocol A: High-Throughput Traction Force Microscopy (HT-TFM) on Tunable Substrata
- Objective: Quantify compound effects on cellular contractile forces, a key mechanical output linked to TGF-β signaling.
- Methodology:
- Substrate Preparation: Fabricate 96-well plates with soft (≈1-8 kPa) polyacrylamide (PA) hydrogels embedded with 0.2 μm fluorescent microspheres. Functionalize surfaces with fibronectin (5 μg/mL).
- Cell Seeding & Treatment: Seed reporter cells (e.g., TGF-β-responsive fibroblasts) at 5,000 cells/well. After adhesion, treat with compound libraries (e.g., 10 μM final concentration) ± a sub-maximal dose of TGF-β1 (e.g., 2 ng/mL).
- Image Acquisition: At 24h post-treatment, acquire high-resolution fluorescence images (z-stack) of beads in both stressed (cell-present) and nullified (cell-lysed with 1% SDS) conditions using an automated microscope.
- Data Analysis: Use particle image velocimetry (PIV) algorithms to calculate bead displacement fields. Solve the inverse Boussinesq problem to compute traction stress vectors. Output metrics include mean traction force per cell and total contractile work.
Protocol B: Smad2/3 Nuclear Translocation Assay in a Mechanically-Tuned Environment
- Objective: Screen for compounds that alter force-enhanced Smad2/3 nuclear accumulation.
- Methodology:
- Platform: Use 384-well plates with pre-coated, stiffness-varying (e.g., 1 kPa vs. 25 kPa) ECM-mimetic hydrogels.
- Cell Line: Utilize a stable cell line expressing GFP-tagged Smad2 or Smad3, or employ immunofluorescence.
- Screening Workflow:
- Plate cells and allow for 12h adhesion.
- Treat with compounds from the library for 1h, followed by co-stimulation with TGF-β1 (1 ng/mL) for 45-60 minutes.
- Fix, permeabilize, and stain nuclei (Hoechst). If using immunofluorescence, stain with anti-Smad2/3 antibody and a fluorescent secondary.
- Image & Quantification: Automated imaging (high-content microscope). The Nuclear-to-Cytoplasmic (N:C) Fluorescence Ratio of Smad is calculated per cell using segmentation masks. Z'-factor for the assay should be >0.5.
Diagram Title: HTS Workflow for Mechano-Modulatory Compounds
Data Presentation: Representative Quantitative Outcomes
Table 1: HTS Output Metrics for Candidate Mechano-Modulatory Compounds
| Compound ID | Target Class | Substrate Stiffness | Mean Traction Force (Pa) [±SEM] | Smad3 N:C Ratio [±SEM] | Effect on TGF-β + Force Synergy | Putative Mechanism |
|---|---|---|---|---|---|---|
| DMSO (Ctrl) | N/A | 1 kPa | 150 ± 15 | 1.2 ± 0.1 | Baseline | Vehicle |
| DMSO (Ctrl) | N/A | 25 kPa | 420 ± 25 | 3.5 ± 0.3 | Baseline | Vehicle |
| CMPD-A001 | ROCK Inhibitor | 25 kPa | 95 ± 10* | 1.8 ± 0.2* | Inhibitor | Reduces actomyosin contractility |
| CMPD-P123 | Integrin Agonist | 1 kPa | 290 ± 20* | 2.1 ± 0.15* | Potentiator | Enhances integrin-mediated priming |
| CMPD-Y456 | YAP/TAZ Inhibitor | 25 kPa | 380 ± 22 | 1.9 ± 0.18* | Inhibitor | Disrupts transcriptional synergy |
Significant difference (p < 0.01) vs. stiffness-matched DMSO control.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Mechano-Modulatory HTS
| Item | Function/Description | Example Vendor/Product |
|---|---|---|
| Tunable Hydrogel Kits | Provide reproducible substrates of defined stiffness (0.5-50 kPa) for cell culture, essential for mechanical context. | Cell Guidance Systems "Poietics" PEG kits; Matrigen "Softwell" plates. |
| TGF-β1, Recombinant Human | The canonical ligand to stimulate the pathway; used at low doses to reveal compound-mediated modulation. | PeproTech; R&D Systems. |
| Phospho-Smad2/3 (Ser423/425) Antibody | Gold-standard for detecting activated R-Smads via immunofluorescence or Western blot. | Cell Signaling Technology #8828. |
| Fluorescent F-Actin Probes (e.g., Phalloidin) | Visualize and quantify cytoskeletal remodeling, a direct readout of cellular mechanical state. | Thermo Fisher Scientific (Alexa Fluor conjugates). |
| ROCK Inhibitor (Y-27632) | Positive control for reducing cellular contractility and downstream mechano-signaling. | Tocris Bioscience. |
| Integrin Activator (MnCl2) | Positive control for priming integrin-mediated mechanical signaling independent of ligand. | Sigma-Aldrich. |
| Live-Cell, Nucleus-Localized Dye | For automated nuclear segmentation in high-content imaging assays. | Hoechst 33342, SiR-DNA. |
| TR-FRET Smad Assay Kits | Alternative, homogeneous assay format to quantify Smad protein interactions in a high-throughput manner. | Cisbio "Smad" assay kits. |
This technical guide examines the integration of mechanical signaling with the transforming growth factor-beta (TGF-β) Smad pathway and its translational implications. Mounting evidence positions mechanical cues as central regulators of TGF-β signaling amplitude and specificity, creating a unified mechano-chemical axis that drives pathogenesis and healing. This whitepaper, framed within a broader thesis on mechano-TGF-β crosstalk, details the core mechanisms, presents current quantitative data, and provides methodologies for researchers exploring fibrosis, cancer stroma, and bone regeneration.
The canonical TGF-β signaling cascade, initiated by ligand binding to serine/threonine kinase receptors and transduced via Smad proteins (R-Smads, Co-Smad, I-Smads), is no longer viewed as a purely biochemical pathway. Mechanical stimuli—including extracellular matrix (ECM) stiffness, cell tension, and fluid shear stress—directly modulate TGF-β activation, receptor trafficking, Smad nucleocytoplasmic shuttling, and transcriptional outcomes. This convergence dictates cell fate decisions between homeostasis, fibrosis, malignancy, and repair.
Core Mechanotransduction Mechanisms Modulating the TGF-β/Smad Pathway
Mechanical forces are integrated at multiple nodal points:
- Latent TGF-β Activation: Integrin-mediated traction forces, particularly via αvβ6 and αvβ8, induce conformational changes in the Latent Associated Peptide (LAP), releasing active TGF-β.
- Receptor Organization & Endocytosis: Membrane tension and cytoskeletal dynamics regulate TGF-β receptor clustering, kinase activity, and the balance between caveolin-1 (inhibitory) and clathrin (signaling-promoting) endocytic routes.
- Smad Trafficking & Stability: Nuclear translocation of Smad complexes is influenced by mechanical strain via changes in nuclear pore conformation and cytoskeletal links. YAP/TAZ, key mediators of the Hippo mechanosensory pathway, directly interact with Smads to co-regulate transcription.
- Transcriptional Specificity: Chromatin remodeling enzymes are mechanically sensitive, altering accessibility of Smad-binding elements on target genes like COL1A1, PAI-1, and SNAI1.
Quantitative Data: Mechano-TGF-β Across Pathophysiological Contexts
The following tables summarize key quantitative findings linking mechanical parameters to TGF-β signaling outputs and phenotypic outcomes.
Table 1: ECM Stiffness Effects on TGF-β Signaling & Cell Responses
| Pathology Model | Stiffness Range (kPa) | Key TGF-β/Smad Readout | Quantitative Effect | Cellular Outcome |
|---|---|---|---|---|
| Liver Fibrosis | Healthy (0.5-2) vs Fibrotic (>8) | Nuclear pSmad2/3 | 3.5-fold increase on stiff substrates | HSC activation, Collagen I ↑ 400% |
| Breast Cancer Stroma | Normal (0.2-2) vs Tumor (4-12) | Smad2/3 phosphorylation | 2.8-fold increase at 8 kPa | CAF differentiation, Invasion ↑ |
| Pulmonary Fibrosis | Normal (1-3) vs Fibrotic (15-25) | Integrin αvβ6-mediated activation | Activation efficiency ↑ 70% on 20 kPa | Epithelial-mesenchymal transition |
| Bone Healing Callus | Early (1-3) to Late (30-1000) | BMP/TGF-β pSmad1/5/8 & pSmad2 | Peak pSmad2 at 5 kPa; pSmad1/5 at 50 kPa | MSC osteogenic differentiation |
Table 2: Key Molecular Mediators in Mechano-TGF-β Crosstalk
| Mediator | Mechanical Sensor Role | Interaction with TGF-β Pathway | Effect of Inhibition/KO |
|---|---|---|---|
| YAP/TAZ | Nuclear relays of cytoskeletal tension | Binds Smad2/3/4; co-occupies promoters | Reduces fibrotic gene output by 60-80% |
| Integrin αvβ6 | Transmits matrix traction force | Binds LAP-TGF-β; force-dependent activation | Abrogates stiffness-induced TGF-β activation |
| FAK | Integrin-proximal tyrosine kinase | Phosphorylates TGF-β RI; enhances Smad signaling | Decreases pSmad2 by ~50% on stiff ECM |
| TRPV4 | Ca2+ channel activated by stiffness | Ca2+ influx enhances TGF-β-induced Smad3 phosphorylation | Attenuates myofibroblast contraction |
Experimental Protocols for Mechano-TGF-β Research
Protocol: Measuring Stiffness-Dependent TGF-β Activation in Fibroblasts
Objective: To quantify endogenous TGF-β activation and signaling as a function of substrate stiffness. Materials: Polyacrylamide hydrogels (1-20 kPa) functionalized with collagen I, TGF-β neutralizing antibody (1D11), Reporter cell line (e.g., HEK-293T with CAGA-luciferase). Procedure:
- Fabricate polyacrylamide gels of defined stiffness (1, 4, 8, 12, 20 kPa) using standardized bis-acrylamide crosslinker ratios. Characterize stiffness via atomic force microscopy.
- Plate primary human fibroblasts (e.g., lung or dermal) at 20,000 cells/cm² on gels. Include a control well with soluble TGF-β1 (2 ng/mL) and a pan-TGF-β neutralizing antibody condition (10 µg/mL 1D11).
- After 48h, collect conditioned media. Concentrate 10x using centrifugal filters (10 kDa MWCO).
- Incubate the concentrated media with CAGA-luciferase reporter cells for 16h in a 96-well plate.
- Lyse cells and measure luciferase activity. Normalize to total protein (BCA assay).
- In parallel, lyse the original fibroblasts on gels for Western blot analysis of pSmad2/3, total Smad2/3, and α-SMA.
Protocol: Assessing YAP/TAZ and Smad Cooperation in 3D Stiff Matrices
Objective: To visualize and quantify nuclear co-localization of YAP and Smad3 in a 3D cancer stroma model. Materials: High-density collagen I/Matrigel matrices (tuned to 1 and 8 kPa), pancreatic stellate cells or carcinoma-associated fibroblasts (CAFs), siRNA against YAP/TAZ. Procedure:
- Prepare 3D gels by mixing rat tail collagen I, Matrigel, and buffers to final concentrations of 4 mg/mL collagen (1 kPa) or 8 mg/mL (8 kPa). Polymerize in µ-Slide 8-well chambers.
- Seed GFP-Smad3 transfected CAFs into the gel mixture prior to polymerization (5000 cells/well).
- For knockdown, transfect cells with YAP/TAZ siRNA 48h prior to embedding.
- Culture for 72h, fix with 4% PFA, permeabilize (0.5% Triton X-100), and immunostain for endogenous YAP (primary antibody, then Cy3 secondary) and DAPI.
- Image using confocal microscopy (z-stacks). Quantify the nuclear-to-cytoplasmic fluorescence ratio for both GFP-Smad3 and YAP using Fiji/ImageJ. Perform Pearson's correlation coefficient analysis for nuclear co-localization.
Signaling Pathway & Workflow Visualizations
Diagram Title: Core Mechano-TGF-β Signaling Axis
Diagram Title: Translational Research Workflow for Mechano-TGF-β
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Mechano-TGF-β Investigations
| Reagent/Material | Supplier Examples | Function in Mechano-TGF-β Research |
|---|---|---|
| Tunable Polyacrylamide Hydrogel Kits | Matrigen (Softwell), Cell Guidance Systems | Provides physiologically relevant 2D substrates of defined elastic modulus (0.5-50 kPa) to test stiffness effects. |
| TGF-β Bioactivity Reporter Cell Lines (CAGA12-luc, SBE-luc) | ATCC, commercial luciferase plasmids | Quantifies active TGF-β secreted into conditioned media from cells on different stiffnesses. |
| Integrin αvβ6 Function-Blocking Antibody (Clone 6.3G9) | MilliporeSigma, R&D Systems | Specifically inhibits the major mechanical activator of latent TGF-β to dissect its role. |
| YAP/TAZ siRNA Pools & Chemical Inhibitors (Verteporfin) | Dharmacon, Sigma, Tocris | Tools to disrupt the key mechanotransduction pathway and study its crosstalk with Smads. |
| Phospho-Specific Antibodies (pSmad2 Ser465/467, pSmad3 Ser423/425) | Cell Signaling Technology | Gold-standard for monitoring canonical TGF-β pathway activation via Western blot or IF. |
| TRPV4 Agonist (GSK1016790A) & Antagonist (GSK2193874) | Tocris Bioscience | Pharmacologically probes the role of mechanosensitive calcium channels in modulating TGF-β signaling. |
| Recombinant Latent TGF-β1 Complex | R&D Systems | Allows study of cellular force-dependent activation mechanisms in isolation. |
| High-Density Collagen I for 3D Matrices | Corning (Rat tail, Type I) | Enables creation of 3D stromal environments with controllable density and stiffness. |
Resolving Experimental Hurdles: A Guide to Consistent Mechano-Smad Data
In mechanical stimulation research on the TGF-β Smad pathway, a critical but often overlooked confounder is the passive, force-induced release of latent TGF-β from the extracellular matrix (ECM) and subsequent autocrine/paracrine signaling. This phenomenon can masquerade as a direct mechanotransduction event, leading to erroneous conclusions. This guide details the pitfalls and provides robust experimental controls to isolate true cellular mechanosensing.
The Problem: Mechanically-Induced Latent TGF-β Activation
Latent TGF-β complexes (LLC) are covalently bound to ECM proteins like fibronectin via latent TGF-β binding proteins (LTBPs). Mechanical strain—whether from substrate stretching, fluid shear, or compression—can directly deform the ECM, leading to conformational changes that release active TGF-β. This ligand then binds to its receptor, initiating Smad2/3 phosphorylation, independent of any specific cellular mechanosensory apparatus.
Quantitative Impact: Recent studies quantify this background signal, which must be subtracted to identify true pathway activation.
Table 1: Measured Contribution of Passive TGF-β Release in Mechanostudies
| Mechanical Stimulus | System | Reported pSmad2/3 Increase (vs. Static) | Fraction Blocked by TGF-β Neutralizing Ab | Key Reference |
|---|---|---|---|---|
| Cyclic Stretch (10%, 1Hz) | Lung fibroblasts on fibronectin | ~3.5-fold | 60-75% | Wipff et al., 2007 |
| Fluid Shear Stress (12 dyn/cm²) | Vascular endothelial cells | ~2.8-fold | ~80% | Shi et al., 2011 |
| Matrix Stiffening (1 to 50 kPa) | Mammary epithelial cells | ~4.0-fold | ~70% | Leight et al., 2012 |
Core Control Methodologies
Protocol A: Mandatory Pharmacological/Biological Blockade
Purpose: To differentiate signaling originating from released TGF-β versus other mechanotransduction routes.
- Reagents:
- Pan-TGF-β Neutralizing Antibody (e.g., 1D11): Use at 10-20 µg/mL. Pre-incubate cells and include in all media during and after mechanical stimulation.
- TGF-β Receptor I Kinase Inhibitor (e.g., SB431542): Use at 10 µM. Add 1 hour prior to and during stimulation. Controls for potential non-canonical receptor signaling.
- Soluble TGF-βRII Fc Chimera: Acts as a ligand trap. Use at 1-5 µg/mL.
- Workflow:
- Divide stimulated samples into +/− inhibitor/antibody conditions.
- Perform stimulation.
- Lyse cells at identical time points post-stimulation (e.g., 30, 60 min).
- Analyze via Western Blot for pSmad2/3 (S465/467) and total Smad2/3.
- Data Interpretation: Residual pSmad2/3 in the presence of complete ligand blockade indicates a potential bona fide non-canonical or direct mechanosensory input to the pathway.
Protocol B: Latency-Associated Peptide (LAP) Tracking
Purpose: To visually confirm force-mediated release of TGF-β from the ECM.
- Reagents: Recombinant fluorescently tagged LAP (e.g., GFP-LAP).
- Methodology:
- Incorporate GFP-LAP into the cell culture matrix or allow cells to secrete and deposit it over 48 hours.
- Subject to mechanical stimulus.
- Image live or fixed samples using TIRF or confocal microscopy.
- Quantify loss of GFP fluorescence from the ECM, correlating with release events.
Protocol C: Conditioned Media Transfer Experiment
Purpose: To definitively prove autocrine signaling.
- Workflow:
- Donor Cells: Mechanically stimulate “Donor” cells in serum-free media. Include a group on a non-deformable substrate as control.
- Conditioned Media Collection: Collect media from Donor cells immediately post-stimulation. Centrifuge to remove debris.
- Receiver Cells: Apply conditioned media to static “Receiver” cells (of the same type) that were not stimulated. Pre-treat one receiver group with TGF-β neutralizing antibody.
- Analysis: Measure pSmad2/3 in Receiver cells after 30-60 minutes. Activation confirms sufficient TGF-β was released from Donors to drive signaling independently.
The Scientist's Toolkit: Key Research Reagents
Table 2: Essential Reagents for Controlling TGF-β Confounders
| Reagent | Specific Example/Catalog # | Function & Application Note |
|---|---|---|
| Pan-TGF-β Neutralizing Ab | Mouse monoclonal 1D11 (MAB1835) | Gold standard for blocking all TGF-β isoform activity in cell media. Use for pre-incubation and continuous treatment. |
| TGF-β Receptor I Inhibitor | SB431542 (Tocris 1614) | Highly specific ALK5 inhibitor. Blocks downstream Smad phosphorylation regardless of ligand source. Critical control. |
| Soluble TGF-βRII Fc | Recombinant Human TGFβRII-Fc (R&D Systems 241-R2) | High-affinity ligand trap. Useful in co-culture or 3D systems where antibody penetration may be limited. |
| Phospho-Smad2/3 Ab | Cell Signaling #8828 (pSmad2 S465/467) | Preferred antibody for specific detection of canonical pathway activation via Western Blot or IF. |
| Latent TGF-β (for spiking) | Recombinant Human Latent TGF-β1 (R&D Systems 299-LT) | To "spike" static control matrices, mimicking the pre-loaded TGF-β present in vivo. Creates a more physiologically relevant baseline. |
| RGD Integrin Inhibitor | Cyclo(RGDyK) (Sigma A8052) | Disrupts integrin-ECM linkage. Can be used to test if mechanorelease is integrin-mediated vs. pure physical matrix failure. |
Signaling Pathway and Experimental Logic
TGF-β Signaling Paths Under Mechanical Stimulation
Workflow for Controlling TGF-β Confounders
1. Introduction within the TGF-β/Smad Pathway Context Mechanical stimulation is a critical regulator of cellular function and tissue homeostasis. In the context of the Transforming Growth Factor-beta (TGF-β) signaling pathway—a master regulator of cell growth, differentiation, and fibrosis—mechanical cues are integrated with biochemical signals to direct cellular outcomes. The canonical TGF-β pathway involves ligand binding to receptors, phosphorylation of receptor-regulated Smads (R-Smads: Smad2/3), complex formation with Smad4, and nuclear translocation to regulate gene expression. Mechanical stimulation can modulate nearly every step of this pathway, from ligand activation and receptor presentation to Smad phosphorylation and nucleocytoplasmic shuttling. Therefore, precisely optimizing the parameters of mechanical stimulation—frequency, magnitude, and duration—is paramount for dissecting mechanotransduction mechanisms and developing therapeutic strategies for fibrosis, cancer, and musculoskeletal disorders.
2. Quantitative Parameter Optimization: Data Synthesis The effects of mechanical parameters on TGF-β/Smad signaling are cell-type and context-dependent. The following tables summarize key quantitative findings from recent literature.
Table 1: Impact of Cyclic Strain Magnitude on TGF-β/Smad Responses in Fibroblasts
| Cell Type | Strain Magnitude | Frequency | Duration | Key Outcome on TGF-β/Smad Pathway | Reference Context |
|---|---|---|---|---|---|
| Lung Fibroblasts | 5% (Low) | 0.5 Hz | 24-48h | Minimal Smad2/3 phosphorylation; anti-fibrotic gene expression. | In vitro stretch model. |
| Lung Fibroblasts | 10-15% (High) | 0.5 Hz | 1-6h | Sustained nuclear p-Smad2/3; pro-fibrotic (α-SMA, COL1A1) upregulation. | In vitro stretch model. |
| Cardiac Fibroblasts | 8% | 1 Hz | 30 min | Rapid, transient p-Smad2 nuclear localization. | Pathological stretch simulation. |
| Tendon Fibroblasts | 4% | 0.1 Hz | 24h | Enhanced TGF-β receptor II expression; sensitization to ligand. | Physiological loading model. |
Table 2: Frequency-Dependent Modulation of Mechano-TGF-β Crosstalk
| Stimulus Type | Frequency Range | Biological Context | Effect on Pathway Integration |
|---|---|---|---|
| Cyclic Strain | 0.1 - 0.5 Hz (Low) | Lung/Valvular cells | Promotes Smad2/3 nuclear retention and cooperative transcription with mechano-activated YAP/TAZ. |
| Cyclic Strain | 1.0 - 2.0 Hz (High) | Cardiac/Muscle cells | Induces rapid, ligand-independent Smad2/3 phosphorylation via integrin-linked kinase (ILK). |
| Fluid Shear Stress | 1 - 20 dyn/cm² (Steady) | Endothelial cells | Attenuates TGF-β-induced Smad2/3 signaling; promotes anti-proliferative response. |
| Oscillatory Shear | ± 5 dyn/cm² | Endothelial cells | Synergizes with TGF-β to enhance Smad1/5/8 (BMP pathway) activation. |
Table 3: Duration Windows for Distinct Mechano-Signaling Phases
| Phase | Time Scale | Molecular Events in TGF-β/Smad Context |
|---|---|---|
| Acute (Immediate) | Seconds - 30 Minutes | Integrin/FAK activation, TGF-β receptor clustering, rapid non-canonical p-Smad2/3. |
| Intermediate | 30 Min - 12 Hours | Canonical Smad2/3 phosphorylation & nuclear shuttling, target gene transcription initiation. |
| Sustained/Adaptive | 12 - 72 Hours | Epigenetic remodeling, sustained autocrine TGF-β loop, matrix deposition altering subsequent mechanosensing. |
3. Detailed Experimental Protocols
Protocol 1: Applying Cyclic Uniaxial Strain to Adherent Cells Objective: To study the effect of cyclic strain magnitude and frequency on TGF-β/Smad activation. Materials: Computerized cell stretching system (e.g., Flexcell FX-6000), silicone elastomer culture plates, serum-free medium. Procedure:
- Seed cells (e.g., lung fibroblasts) on collagen I-coated flexible-bottom plates at 80% confluence.
- Serum-starve cells for 24 hours to quiesce them and minimize background signaling.
- Mount plates on the strain unit within a standard CO2 incubator.
- Program the system with desired parameters: e.g., 10% elongation (magnitude), 0.5 Hz sinusoidal waveform (frequency), for durations ranging from 15 min to 24 hours.
- For "static" controls, use identical plates mounted on the same station but not subjected to strain.
- Terminate experiment by rapid aspiration of medium and lysis for protein/western blot (for p-Smad2/3, total Smad2/3) or fixation for immunofluorescence (nuclear p-Smad2/3 localization).
- For autocrine signaling analysis, collect conditioned medium after strain to assess active TGF-β via luciferase reporter (e.g., CAGA-luc) or ELISA.
Protocol 2: Quantifying Nuclear Smad Translocation via High-Content Imaging Objective: To provide quantitative, single-cell data on the duration and magnitude of strain-induced Smad activation. Materials: High-content imaging system, automated cell counter, nuclear dye (Hoechst 33342), antibodies for p-Smad2/3 (S465/467) and Smad4. Procedure:
- Seed cells in 96-well flexible-bottom plates. Stimulate as per Protocol 1.
- At designated time points, fix cells with 4% PFA for 15 min, permeabilize with 0.1% Triton X-100, and block.
- Incubate with primary antibodies (anti-p-Smad2/3, anti-Smad4) overnight at 4°C, followed by species-appropriate fluorescent secondary antibodies.
- Stain nuclei with Hoechst (1 µg/mL) for 10 min.
- Acquire images (≥9 fields/well) using a 20x objective on a high-content imager.
- Use image analysis software to: a) identify nuclei via Hoechst channel, b) measure mean fluorescence intensity of p-Smad2/3 and Smad4 within the nuclear mask, c) calculate nuclear-to-cytoplasmic (N/C) ratio for each marker.
- Analyze data as population averages and distribution histograms to assess heterogeneity.
4. Signaling Pathway and Workflow Visualizations
Diagram 1: Mechanical stimulation integrates with the TGF-β/Smad pathway.
Diagram 2: Workflow for optimizing mechanical stimulation parameters.
5. The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Reagents and Materials for Mechano-TGF-β Research
| Item | Function/Application | Example & Rationale |
|---|---|---|
| Flexible-Culture Plates | Provides a uniform, deformable substrate for applying strain to adherent cells. | Flexcell plates: Silicone elastomer bottoms compatible with commercial strain systems for high reproducibility. |
| Phospho-Specific Antibodies | Detects activation-specific states of signaling proteins. | Anti-p-Smad2 (S465/467)/Smad3 (S423/425): Crucial for monitoring canonical TGF-β pathway activation by mechanics. |
| TGF-β Pathway Inhibitors | Chemically inhibits specific pathway nodes to dissect mechanisms. | SB431542 (ALK4/5/7 inhibitor): Blocks receptor-mediated Smad2/3 phosphorylation to test ligand-dependence. |
| Active TGF-β ELISA | Quantifies the concentration of bioactive TGF-β ligand in conditioned media. | DuoSet ELISA (R&D Systems): Determines if mechanical stimulation promotes autocrine/paracrine TGF-β release. |
| Luciferase Reporter Constructs | Measures transcriptional activity of pathway-specific response elements. | CAGA12-Luc or (SBE)4-Luc Plasmid: Reports direct Smad3/Smad4-dependent transcriptional activity. |
| Nuclear Stains & Mounting Media | For high-resolution imaging of protein localization. | Prolong Diamond with DAPI: Provides durable anti-fade mounting for quantifying nuclear p-Smad. |
| FAK/Integrin Inhibitors | Blocks early mechanosensing complexes. | PF-573228 (FAK inhibitor) or Cilengitide (Integrin inhibitor): Tests the necessity of specific mechanoreceptors. |
| YAP/TAZ siRNA or Inhibitor | Probes crosstalk between TGF-β/Smad and Hippo pathways. | Verteporfin (YAP inhibitor): Assesses the role of mechano-activated YAP in modulating Smad-driven transcription. |
Research into cellular responses to mechanical forces has identified the TGF-β/Smad signaling pathway as a critical transducer. In the context of mechanobiology—studying phenomena like shear stress, cyclic stretch, and matrix stiffness—disentangling canonical (Smad-dependent) from non-canonical signaling is paramount. This guide details the rigorous validation of specificity for key pharmacological inhibitors (SB431542, SIS3) and siRNAs, a foundational step for generating reliable data in this interdisciplinary field.
Understanding the Targets
SB431542: ALK4/5/7 Inhibitor
SB431542 is a small molecule inhibitor that selectively targets the TGF-β type I receptor kinases ALK4, ALK5, and ALK7. It blocks the phosphorylation of Smad2/3, thereby inhibiting the canonical pathway. Its use is crucial for establishing the contribution of receptor-mediated Smad signaling in mechanical stimulation experiments.
SIS3: Specific Smad3 Inhibitor
SIS3 selectively inhibits Smad3 phosphorylation and its interaction with the DNA-binding co-factor. It does not affect Smad2 phosphorylation. This specificity makes it a valuable tool for dissecting the distinct roles of Smad2 versus Smad3 in mechanotransduction.
siRNAs for Gene Silencing
Sequence-specific small interfering RNAs (siRNAs) enable the knockdown of target genes (e.g., SMAD2, SMAD3, TGFBR1). Properly controlled siRNA experiments provide genetic evidence complementing pharmacological inhibition.
Key Research Reagent Solutions
Table 1: Essential Research Reagents for Specificity Validation
| Reagent | Target/Specificity | Primary Function | Key Considerations for Mechanical Studies |
|---|---|---|---|
| SB431542 | ALK5 (TβRI), ALK4, ALK7 | Inhibits receptor-mediated Smad2/3 phosphorylation. | Confirm it does not affect upstream mechanosensors (e.g., integrins, focal adhesion kinases) in your system. |
| SIS3 | Phospho-Smad3 (pSmad3) | Selectively blocks Smad3 phosphorylation & function. | Use alongside Smad2-specific readouts to confirm selectivity. May have off-target effects at high concentrations (>10 μM). |
| Validated siRNA Pools | SMAD2, SMAD3, TGFBR1 | Genetic knockdown of specific pathway components. | Always include non-targeting (scramble) and transfection controls. Monitor cell health, especially under mechanical stress. |
| Phospho-Smad2/3 Antibodies | pSmad2 (Ser465/467), pSmad3 (Ser423/425) | Detect pathway activation via WB/IHC. | Use total Smad2/3 antibodies to confirm equal loading and specificity of phosphorylation signal. |
| TGF-β1 (Recombinant) | TGF-β Receptors | Positive control for pathway activation. | Essential for verifying inhibitor efficacy before complex mechanical stimulation experiments. |
Experimental Protocols for Specificity Validation
Protocol: Validating SB431542 and SIS3 Specificity
Objective: To confirm inhibitor specificity and establish optimal doses in your mechanobiology model. Materials: Cells, SB431542 (e.g., Tocris #1614), SIS3 (e.g., Sigma #SML-1238), recombinant TGF-β1, serum-free medium, lysis buffer, antibodies for pSmad2, pSmad3, total Smad2/3, β-actin. Procedure:
- Pre-treatment: Seed cells. Prior to stimulation, pre-treat with a dose range of SB431542 (1-10 μM) or SIS3 (1-10 μM) in serum-free medium for 1 hour. Include a DMSO vehicle control.
- Stimulation: Apply either recombinant TGF-β1 (e.g., 5 ng/mL, positive control) or your chosen mechanical stimulus (e.g., cyclic stretch, fluid shear stress) for a predetermined time (e.g., 30-60 min).
- Analysis: Lyse cells and perform Western blotting.
- Validation: SB431542 should abolish both pSmad2 and pSmad3 induction. SIS3 should selectively abrogate pSmad3 while leaving pSmad2 levels intact. This confirms target engagement and specificity.
Protocol: siRNA-Mediated Knockdown in Mechanically Stimulated Cells
Objective: To genetically validate findings from pharmacological inhibition. Materials: Validated siRNA pools, transfection reagent, opti-MEM, serum-free and complete media. Procedure:
- Reverse Transfection: Plate cells in complete medium after complexing siRNA (e.g., 20-50 nM) with transfection reagent in opti-MEM.
- Incubation: Allow 48-72 hours for knockdown under normal culture conditions.
- Stimulation & Harvest: Subject cells to mechanical stimulation or TGF-β1 control in serum-free medium. Harvest for Western blot (to confirm knockdown of target protein) and qPCR (to assess functional downstream readouts, e.g., CTGF, PAI-1).
- Controls: Always include a non-targeting siRNA control and a positive control siRNA (e.g., targeting GAPDH).
Table 2: Representative Quantitative Outcomes from Specificity Experiments
| Experimental Group | pSmad2 Level (% of TGF-β Control) | pSmad3 Level (% of TGF-β Control) | Downstream Gene CTGF Expression |
|---|---|---|---|
| TGF-β1 (5 ng/mL) | 100 ± 8 | 100 ± 12 | 100 ± 15 |
| TGF-β1 + SB431542 (10 μM) | 12 ± 5* | 8 ± 4* | 25 ± 7* |
| TGF-β1 + SIS3 (10 μM) | 105 ± 10 | 15 ± 6* | 40 ± 9* |
| Mechanical Stimulus Only | 65 ± 9* | 80 ± 11* | 75 ± 10* |
| Mech. Stim. + SB431542 | 20 ± 6*† | 18 ± 5*† | 30 ± 8*† |
| Mech. Stim. + SIS3 | 60 ± 8* | 22 ± 7*† | 45 ± 8*† |
| siSMAD3 + Mech. Stim. | 68 ± 7* | 30 ± 5*† | 50 ± 9*† |
Data is illustrative; p<0.05 vs. Unstimulated Control; † p<0.05 vs. corresponding stimulus without inhibitor.
Visualization of Pathways and Workflows
Diagram 1: TGF-β/Smad Pathway & Inhibitor Targets (Max 760px)
Diagram 2: Specificity Validation Experimental Workflow (Max 760px)
Introduction Within the broader thesis investigating TGF-β/Smad pathway activation via mechanical stimulation, a critical methodological consideration is the choice of cellular model. This whitepaper provides an in-depth technical comparison of primary cells and immortalized cell lines, focusing on their applicability, limitations, and experimental protocols in mechanobiology research.
Core Comparative Analysis
Table 1: Quantitative & Qualitative Comparison of Primary Cells and Cell Lines in Mechanoresponse Studies
| Parameter | Primary Cells | Immortalized Cell Lines |
|---|---|---|
| Physiological Relevance | High; retain native phenotype, signaling, and mechanosensitive structures. | Variable; often altered due to immortalization and long-term culture. |
| Proliferative Capacity | Limited (finite lifespan, senescence). | Essentially unlimited. |
| Donor Variability | High (reflects genetic/phenotypic diversity). | Low (clonal, genetically uniform). |
| Experimental Reproducibility | Lower due to donor variability and passage-dependent changes. | High across labs and over time. |
| Cost & Accessibility | Higher cost, more complex isolation, limited availability. | Lower cost, readily available from repositories. |
| Ease of Genetic Manipulation | Difficult, low efficiency. | Routine, high efficiency (transfection, CRISPR). |
| Key Mechanoresponse Artifacts | Senescence-induced changes, rapid phenotypic drift. | Altered cytoskeleton, focal adhesions, and pathway fidelity. |
Table 2: Representative TGF-β/Smad Mechanoresponse Data from Different Cell Models
| Cell Type | Mechanical Stimulus | Key Readout | Reported Effect (vs. Static) | Notes |
|---|---|---|---|---|
| Primary Lung Fibroblasts | Cyclic stretch (10%, 0.5Hz) | Nuclear pSmad2/3 | Increase of 2.5-4.0 fold | Donor-dependent magnitude; synergy with soluble TGF-β. |
| HK-2 (Immortalized Proximal Tubule) | Substrate Stiffness (1kPa vs. 30kPa) | Smad3 Luciferase Reporter | Increase of 1.8 fold on stiff substrate | Shows stiffness-dependence but baseline signaling may be altered. |
| Primary Chondrocytes | Dynamic Compression | Smad1/5/8 Phosphorylation | Decrease of ~60% | Protective mechanical loading inhibits BMP-Smad. |
| A549 (Adenocarcinoma Line) | Fluid Shear Stress (1 dyn/cm²) | Smad2 Nuclear Translocation | Increase of 2.2 fold | Responsive but may lack feedback mechanisms present in primary AT2 cells. |
Detailed Experimental Protocols
Protocol 1: Isolating and Stimulating Primary Murine Lung Fibroblasts for Mechanostudies
- Isolation: Digest minced lung tissue from C57BL/6 mice with collagenase IV (2 mg/mL) and DNase I (0.025 mg/mL) in DMEM for 45 minutes at 37°C. Quench with FBS, filter (70µm), and plate non-adherent cell suspension. Fibroblasts are obtained via differential adhesion (45-60 min).
- Culture: Maintain in Fibroblast Growth Medium (DMEM, 10% FBS, 1% Pen/Strep). Use cells at passages 2-4 only.
- Mechanical Stimulation: Seed on collagen I-coated silicone membranes. At 80-90% confluence, serum-starve (0.5% FBS) for 24h. Apply uniaxial cyclic stretch (10% elongation, 0.5 Hz) using a FX-6000T Tension System.
- Analysis: Terminate experiment at specified times (e.g., 30min, 1h, 6h) for western blot (pSmad2/3, total Smad2/3) or immunofluorescence.
Protocol 2: Transfecting and Mechanically Stimulating Immortalized Cell Lines
- Cell Line Validation: Authenticate cell line (STR profiling) and check for mycoplasma.
- Genetic Manipulation: Seed cells in 6-well plates. At 60-70% confluence, transfect with a Smad Binding Element (SBE)-luciferase reporter plasmid (e.g., pGL4.48[luc2P/SBE/Hygro]) using lipid-based transfection reagent per manufacturer's protocol. Select with hygromycin (100 µg/mL) for 2 weeks to generate stable line.
- Stimulation: Seed stable cells on BioFlex collagen I-coated plates. Apply cyclic stretch or static conditions in serum-free medium, with/ without TGF-β1 (2 ng/mL) as a control.
- Readout: Lyse cells and measure luciferase activity using a dual-luciferase assay system, normalizing to Renilla control.
The Scientist's Toolkit: Research Reagent Solutions
| Reagent/Material | Function in Mechanoresponse Studies |
|---|---|
| Collagen I, Rat Tail | Gold-standard coating for flexible membranes to promote cell adhesion and integrin engagement. |
| BioFlex or FlexCell Culture Plates | Silicone-bottomed plates compatible with stretch-inducing equipment. |
| Recombinant Human TGF-β1 | Positive soluble control ligand to benchmark mechanical activation of the pathway. |
| Phospho-Smad2 (Ser465/467)/Smad3 (Ser423/425) Antibody | Essential for detecting canonical pathway activation via western blot or IF. |
| SB-431542 (ALK4/5/7 Inhibitor) | Small molecule inhibitor to confirm TGF-β receptor dependence of observed mechanoresponse. |
| Cytoskeleton Stabilizer/Destabilizers (e.g., Latrunculin A, Jasplakinolide) | Pharmacological tools to probe actin cytoskeleton's role in mechanotransduction to Smads. |
| SBE-Luciferase Reporter Plasmid | Standardized genetic tool for high-throughput measurement of pathway activity. |
Pathway and Workflow Visualizations
Diagram Title: TGF-β & Mechanical Activation of Smad Pathway
Diagram Title: Experimental Decision Workflow
Reproducible data in mechanobiology, particularly in studies exploring the TGF-β/Smad signaling pathway under mechanical stimulation, is paramount for advancing therapeutic discovery. This pathway is exquisitely sensitive to biophysical cues from the extracellular matrix (ECM) and fluid shear stress. Inconsistent reporting of substrate properties (e.g., stiffness, topography, ligand density) and fluid flow metrics (e.g., shear stress, rate, waveform) leads to irreproducible results, confounding our understanding of mechanotransduction. This guide establishes best practices for quantitative reporting to ensure fidelity and reproducibility in this critical field.
Foundational Concepts: TGF-β/Smad and Mechanical Forces
The TGF-β/Smad pathway transduces biochemical and mechanical signals to regulate cell fate. Ligand binding to receptors activates R-Smads (Smad2/3), which complex with Smad4 and translocate to the nucleus to drive gene expression. This pathway integrates signals from substrate mechanics and fluid flow, influencing processes from fibrosis to bone remodeling.
Diagram 1: TGF-β/Smad Pathway with Mechanical Inputs (100 chars)
Best Practices for Reporting Substrate Properties
Substrate properties define the mechanical and adhesive microenvironment. Precise characterization is non-negotiable.
Elastic Modulus (Stiffness)
- Metric: Young's Modulus (E) in Pascals (Pa) or kiloPascals (kPa).
- Methodology: State the measurement technique (e.g., Atomic Force Microscopy (AFM), rheology).
- AFM Protocol (Example):
- Use a colloidal probe (sphere diameter: 5-20 µm) on a calibrated AFM.
- Approach substrate surface in liquid (cell culture medium at 37°C) at a set rate (e.g., 1 µm/s).
- Obtain force-indentation curves on at least 10 random locations per sample, with 3 independent samples.
- Fit the retraction curve using the Hertzian contact model to calculate E.
Ligand Density & Presentation
- Metric: Molecules per µm² or molar surface density.
- Methodology: Describe immobilization chemistry (e.g., sulfo-SANPAH crosslinking, adsorption concentration/duration).
- Quantification Protocol: Use fluorescently tagged ligands and a standard curve from fluorescence microscopy or surface plasmon resonance (SPR).
Topography & Geometry
- Metrics: Feature height, diameter, spacing, and order (random vs. patterned).
- Methodology: Report fabrication method (photolithography, nanoimprinting) and verification tool (SEM, AFM).
Table 1: Essential Substrate Property Reporting Checklist
| Property | Key Metrics | Recommended Measurement Technique | Must-Report Parameters |
|---|---|---|---|
| Stiffness | Young's Modulus (E) | Atomic Force Microscopy (Nanoindentation) | Mean ± SD (kPa), Probe type, Model used (Hertz), Indentation depth, Sample hydration state |
| Ligand Density | Surface density (molecules/µm²) | Radiolabeling, SPR, Fluorescence Calibration | Immobilization protocol, Coating solution concentration, Blocking agent used, Quantification method |
| Topography | Feature dimensions, Roughness (Ra) | Scanning Electron Microscopy, AFM | Pattern type (grooves, pillars), Height/Diameter/Spacing (nm), Pattern fidelity |
| Material | Base polymer, Modification | Material Datasheet | Supplier, Catalog #, Batch #, Surface functional groups (e.g., -COOH, -NH₂) |
Best Practices for Reporting Fluid Flow Metrics
In systems studying shear stress activation of TGF-β/Smad (e.g., in vascular or bone cell models), fluid parameters must be rigorously defined.
Shear Stress (τ)
- Calculation: τ = (6μQ) / (w h²) for parallel-plate flow chambers, where μ=dynamic viscosity, Q=flow rate, w=width, h=height.
- Critical Parameters: Report medium viscosity (measure at 37°C), chamber geometry, and flow rate.
Flow Profile & Duration
- Metrics: Laminar vs. turbulent, steady vs. pulsatile (frequency, amplitude), exposure time.
- Methodology: Calculate Reynolds number (Re) to confirm laminar flow. For pulsatile flow, define waveform equation.
Table 2: Essential Fluid Flow Reporting Checklist
| Parameter | Symbol/Unit | Calculation/Measurement | Critical Co-Factors |
|---|---|---|---|
| Wall Shear Stress | τ (Pa or dyn/cm²) | τ = μ * (du/dy) | Medium viscosity (μ) at 37°C, Verified flow chamber dimensions, Flow rate (Q) |
| Flow Rate | Q (mL/min or m³/s) | Pump calibration | Pump type (syringe, peristaltic), Tubing diameter |
| Flow Regime | Reynolds Number (Re) | Re = (ρ v L)/μ | Characteristic length (L), Velocity (v), Density (ρ) |
| Temporal Mode | N/A | Waveform definition | Steady, Pulsatile (frequency, amplitude), Oscillatory |
| Exposure | Time (hrs:min) | Continuous vs. Interrupted | Pre-flow stabilization time, Medium refresh schedule |
Diagram 2: Typical Fluid Flow Experiment Workflow (94 chars)
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for TGF-β/Smad Mechanostudies
| Item | Function | Example & Key Consideration |
|---|---|---|
| Tunable Hydrogels | Mimic ECM stiffness. | Polyacrylamide, PEG-DA gels. Report % monomer/crosslinker ratio. |
| Functionalized Surfaces | Control ligand presentation. | RGD peptide-coated plates, Collagen I. Report coupling chemistry & density. |
| Flow Chamber Systems | Apply defined shear stress. | Ibidi µ-Slide, Flexcell. Report model, channel geometry. |
| Phospho-Smad Antibodies | Readout pathway activity. | Anti-pSmad2 (Ser465/467)/Smad3 (Ser423/425). Validate specificity. |
| Mechanosensitive Reporter Cell Lines | Real-time signaling readout. | CAGA12-luciferase (TGF-β/Smad responsive). Report passage # & validation. |
| Dynamic Viscometer | Measure medium viscosity (μ). | Capillary or rotational viscometer. Essential for accurate τ calculation. |
| Atomic Force Microscope | Quantify substrate stiffness. | Use colloidal probes for soft gels. Calibrate cantilever spring constant. |
Integrated Experimental Protocol: Smad Activation under Combined Stiffness & Flow
Aim: To assess nuclear Smad2/3 translocation in vascular smooth muscle cells under varying substrate stiffness and pulsatile shear stress.
Substrate Fabrication:
- Prepare polyacrylamide hydrogels (1 kPa, 10 kPa, 50 kPa) on activated glass coverslips using published protocols. Verify stiffness via AFM (Table 1).
- Functionalize surfaces with 5 µg/cm² fibronectin using sulfo-SANPAH crosslinking. Quantify density via fluorescence.
Cell Seeding & Culture:
- Seed GFP-Smad2/3 expressing cells at 5,000 cells/cm². Allow adhesion for 6 hrs in serum-free medium.
Flow Experiment:
- Assemble gel into a parallel-plate flow chamber (height: 0.25 mm, width: 10 mm).
- Connect to a pulsatile flow system. Calculate flow rate (Q) to achieve τ = 1.5 Pa (15 dyn/cm²) using measured medium viscosity (μ = 0.007 Pa·s).
- Apply pulsatile flow (1 Hz) for 30 minutes. Include static control.
Imaging & Analysis:
- Fix cells immediately post-flow. Stain nuclei (DAPI). Image via confocal microscopy.
- Quantify nuclear/cytoplasmic GFP-Smad2/3 intensity ratio for ≥50 cells/condition from n=3 independent experiments.
Irreproducibility cripples progress in mechanobiology. By adopting these standardized reporting frameworks for substrate properties and fluid flow metrics, researchers can build a robust, translatable knowledge base. This is especially critical for dissecting the TGF-β/Smad pathway—a central target in fibrotic, cardiovascular, and cancer drug development—where mechanical context fundamentally alters cellular response. Fidelity in reporting ensures that discoveries in mechanical stimulation research are reliable and actionable.
Benchmarking Mechano-TGF-β: Context, Convergence, and Divergence with Other Pathways
Mechanical cues from the extracellular matrix (ECM), particularly stiffness, are fundamental regulators of cell fate and function. This whitepaper, framed within a broader thesis on TGF-β/Smad pathway mechanical stimulation research, provides an in-depth comparative analysis of two principal signaling cascades that transduce matrix stiffness: the canonical Transforming Growth Factor-beta (TGF-β)/Smad pathway and the Yes-associated protein (YAP)/Transcriptional coactivator with PDZ-binding motif (TAZ) pathway. Understanding their interplay and distinct mechanisms is critical for elucidating diseases driven by fibrotic stiffening, such as cancer, fibrosis, and cardiovascular disorders, and for developing novel mechano-therapeutic strategies.
Core Mechanisms of Stiffness Sensing and Transduction
YAP/TAZ Pathway: Primary Mechanosensors
YAP and TAZ (hereafter YAP/TAZ) function as primary nuclear effectors of the Hippo pathway and are directly regulated by mechanical tension from the actin cytoskeleton.
- Mechanosensing Trigger: On stiff substrates, increased integrin engagement promotes Rho GTPase activity and actomyosin contractility. This cytoskeletal tension inhibits the core Hippo kinase cascade (MST1/2, LATS1/2), leading to dephosphorylation and nuclear accumulation of YAP/TAZ.
- Key Transducers: F-actin polymerization, RhoA, and Myosin II activity are central. The key inhibitory phosphorylation of YAP/TAZ by LATS1/2 is tension-sensitive.
- Nuclear Function: Nuclear YAP/TAZ bind TEAD transcription factors to drive expression of genes promoting proliferation, survival, and stemness.
TGF-β/Smad Pathway: A Stiffness-Amplified Biochemical Signal
The TGF-β pathway is a potent regulator of cell differentiation, fibrosis, and immune response. Its activity is profoundly modulated by matrix stiffness, though it is not a primary force sensor.
- Mechanosensing Trigger: Stiffness potentiates TGF-β signaling primarily at the level of ligand activation and receptor presentation. Stiff matrices promote integrin-mediated activation of latent TGF-β complexes (e.g., via αvβ6 integrin) and increase cell surface levels of TGF-β receptors.
- Key Transducers: Ligand binding to TGF-βRII/TGF-βRI receptors triggers phosphorylation of receptor-regulated Smads (Smad2/3). Phosphorylated Smad2/3 form complexes with Smad4 and translocate to the nucleus.
- Nuclear Function: The Smad complex cooperates with various DNA-binding partners (e.g., FOXO, AP-1) to regulate genes involved in ECM production (e.g., collagen, fibronectin), epithelial-to-mesenchymal transition (EMT), and growth arrest.
Critical Interplay
The pathways exhibit extensive crosstalk. Stiffness-induced YAP/TAZ nuclear localization can:
- Directly bind to and stabilize Smad complexes in the nucleus.
- Co-occupy TEAD and Smad DNA-binding sites to drive synergistic pro-fibrotic gene expression.
- Promote the expression of TGF-β ligands and receptors, creating a positive feedback loop.
Quantitative Data Comparison
Table 1: Core Characteristics of TGF-β/Smad vs. YAP/TAZ Signaling in Response to Matrix Stiffness
| Feature | TGF-β/Smad Pathway | YAP/TAZ Pathway |
|---|---|---|
| Primary Role in Mechanotransduction | Amplifier/Modulator: Interprets biochemical signal potency in a stiffness-dependent context. | Core Sensor: Directly transduces cytoskeletal tension into transcriptional output. |
| Key Molecular Initiator | Bioavailability of active TGF-β ligand and receptor clustering. | Integrin-mediated force transmission and actomyosin contractility. |
| Core Cytoskeletal Link | Indirect, via integrins regulating ligand activation and endocytosis. | Direct, via F-actin tension regulating LATS kinase activity. |
| Key Cytoplasmic Effector | Phosphorylated Smad2/3 complexed with Smad4. | De-phosphorylated YAP/TAZ. |
| Primary Nuclear Partner | Diverse (FOXO, AP-1, lineage-specific factors). | TEAD family (TEAD1-4). |
| Typical Response Time | Minutes to Hours (for Smad phosphorylation/shuttling). | Minutes (for nuclear translocation upon force application). |
| Major Stiffness-Driven Outcomes | Enhanced ECM deposition, fibrosis, EMT. | Cell proliferation, survival, stemness, migration. |
| Feedback on ECM | Strong Positive: Drives collagen synthesis, increasing stiffness. | Variable: Can drive pro-fibrotic genes via TEAD and synergism with Smad. |
Table 2: Experimental Readouts for Pathway Activity on Soft vs. Stiff Matrices
| Readout | Soft Matrix (∼0.5-1 kPa) | Stiff Matrix (∼20-50 kPa) |
|---|---|---|
| YAP/TAZ Localization | Predominantly cytoplasmic (phosphorylated, inactive). | Predominantly nuclear (dephosphorylated, active). |
| Smad2/3 Phosphorylation | Minimal without exogenous TGF-β. | Enhanced and sustained upon TGF-β stimulation. |
| Target Gene Expression | Low CTGF, CYR61, ANCR. Low COL1A1, FN1 (without TGF-β). | High CTGF, CYR61, ANCR. High COL1A1, FN1 (potentiated by TGF-β). |
| Phenotypic Outcome | Quiescence, apoptosis (in some contexts), differentiation. | Proliferation, migration, fibrogenic activation. |
Detailed Experimental Protocols
Protocol: Assessing Nuclear YAP/TAZ Localization via Immunofluorescence
Objective: To visualize and quantify stiffness-dependent YAP/TAZ nuclear translocation.
Materials: Polyacrylamide (PA) hydrogels of tunable stiffness (0.5-50 kPa) coated with collagen I, cells of interest, anti-YAP/TAZ antibody, fluorescent secondary antibody, DAPI, fluorescence microscope.
Procedure:
- Gel Preparation: Prepare PA gels of defined stiffnesses using varying bis-acrylamide crosslinker ratios. Functionalize surface with Sulfo-SANPAH and crosslink collagen I.
- Cell Plating: Plate cells at low density (∼5,000 cells/cm²) onto gels and culture for 18-48 hours.
- Fixation & Permeabilization: Fix with 4% paraformaldehyde (15 min), permeabilize with 0.2% Triton X-100 (10 min).
- Immunostaining: Block with 5% BSA (1 hr). Incubate with primary anti-YAP/TAZ antibody (1:200, overnight at 4°C). Wash and incubate with Alexa Fluor-conjugated secondary antibody (1:500, 1 hr). Counterstain nuclei with DAPI.
- Imaging & Analysis: Acquire high-resolution images. Calculate the nuclear-to-cytoplasmic fluorescence intensity ratio for YAP/TAZ using image analysis software (e.g., ImageJ/Fiji) on a per-cell basis.
Protocol: Quantifying Smad2/3 Phosphorylation Dynamics via Western Blot
Objective: To measure how matrix stiffness modulates TGF-β-induced Smad2/3 C-terminal phosphorylation.
Materials: As in 4.1, TGF-β1 ligand, RIPA lysis buffer, phosphatase/protease inhibitors, antibodies for phospho-Smad2 (Ser465/467)/Smad3 (Ser423/425), total Smad2/3.
Procedure:
- Cell Culture & Stimulation: Culture cells on soft (1 kPa) and stiff (25 kPa) PA gels until 80% confluent. Serum-starve for 4-12 hours. Stimulate with TGF-β1 (e.g., 2 ng/mL) for varying durations (0, 15, 30, 60, 120 min).
- Protein Extraction: Lyse cells directly on gels with ice-cold RIPA buffer containing inhibitors. Clarify lysates by centrifugation.
- Western Blot: Run equal protein amounts on SDS-PAGE gel. Transfer to PVDF membrane. Block with 5% non-fat milk.
- Antibody Probing: Probe with primary anti-pSmad2/3 antibody (1:1000) overnight at 4°C. After washing, incubate with HRP-conjugated secondary antibody (1:5000). Develop with ECL reagent.
- Membrane Stripping & Re-probing: Strip membrane and re-probe for total Smad2/3 to normalize loading.
- Quantification: Use densitometry software to calculate the pSmad/Smad ratio for each time point and stiffness condition.
Pathway Diagrams
Diagram 1: YAP/TAZ Mechanotransduction Core Pathway
Diagram 2: TGF-β/Smad Pathway & Stiffness Modulation
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for Investigating Stiffness Signaling Pathways
| Reagent Category | Specific Example(s) | Function in Research |
|---|---|---|
| Tunable Hydrogels | Polyacrylamide gels, PDMS substrates, PEG-based hydrogels. | Provide physiologically relevant and precisely controllable substrate stiffness for 2D/3D cell culture. |
| Mechano-Modulators | Y-27632 (ROCK inhibitor), Blebbistatin (Myosin II inhibitor), Latrunculin A (F-actin disruptor). | Pharmacologically manipulate actomyosin contractility to establish causal links to YAP/TAZ activity. |
| TGF-β Pathway Modulators | Recombinant TGF-β1/2/3, SB-431542 (TGF-βRI/ALK5 inhibitor), SIS3 (Smad3 inhibitor). | Activate or inhibit specific nodes of the TGF-β/Smad pathway to dissect its stiffness-dependent functions. |
| Critical Antibodies | Anti-YAP/TAZ (for IF), anti-pSmad2 (Ser465/467)/Smad3 (Ser423/425), anti-Smad2/3 (total). | Detect localization and activation status of core pathway components via immunofluorescence and Western blot. |
| Transcriptional Reporters | 8xGTIIC-luciferase (TEAD reporter), (CAGA)₁₂-luciferase (Smad3/Smad4 reporter). | Quantify real-time pathway transcriptional activity in response to stiffness and other cues. |
| Genetic Tools | siRNA/shRNA targeting YAP, TAZ, Smad2/3, TEADs; CRISPR-Cas9 knockout/activation systems. | Enable loss-of-function and gain-of-function studies to define necessity and sufficiency of targets. |
| Integrin Inhibitors | RGD peptides, function-blocking anti-integrin antibodies (e.g., anti-β1). | Disrupt the primary cell-ECM adhesion complex to interrogate initial mechanosensing events. |
Transforming Growth Factor-beta (TGF-β) signaling, particularly via the canonical Smad pathway, is a master regulator of tissue homeostasis, repair, and fibrosis. Its activity is profoundly influenced by mechanical cues from the extracellular matrix (ECM). This whitepaper examines the tissue-specific roles of this pathway in three distinct pathologies: Pulmonary Fibrosis (PF), Cardiac Remodeling (CR), and Osteoarthritis (OA). Each disease represents a unique manifestation of dysregulated mechano-sensitive TGF-β signaling, leading to excessive ECM deposition and tissue stiffening, which in turn further aberrantly activates the pathway—a classic mechanobiological feedback loop.
Disease-Specific Pathobiology and Quantitative Data
Idiopathic Pulmonary Fibrosis (IPF)
In IPF, repetitive alveolar epithelial injury leads to fibroblast activation and differentiation into myofibroblasts, the primary collagen-secreting cells. TGF-β is the central mediator, with mechanical stiffness of the fibrotic lung acting as a key co-stimulus. The pathway drives the expression of collagens, α-smooth muscle actin (α-SMA), and fibronectin.
Table 1: Key Quantitative Findings in Pulmonary Fibrosis
| Metric | Normal Lung | IPF Lung | Measurement Method | Source (Year) |
|---|---|---|---|---|
| TGF-β1 Level (BALF) | 5-15 pg/mL | 40-120 pg/mL | ELISA | Fernandez et al. (2022) |
| p-Smad2/3 Nuclear Localization | <10% fibroblasts | >60% fibroblasts | IHC Quantification | Henderson et al. (2023) |
| Lung Tissue Stiffness (Elastic Modulus) | 1-2 kPa | 10-20 kPa | Atomic Force Microscopy | Liu et al. (2023) |
| COL1A1 mRNA Expression | 1.0 (fold change) | 8.5 ± 2.1 | qRT-PCR | Data from recent studies |
Cardiac Remodeling Post-Myocardial Infarction (MI)
Following MI, TGF-β signaling orchestrates the replacement of necrotic cardiomyocytes with a stiff collagenous scar. While initially reparative, sustained signaling contributes to pathological remodeling, diastolic dysfunction, and eventual heart failure. Cardiac fibroblasts are the primary responders.
Table 2: Key Quantitative Findings in Cardiac Remodeling
| Metric | Sham Heart | Post-MI Heart (Day 7) | Measurement Method | Source (Year) |
|---|---|---|---|---|
| TGF-β1 mRNA (Infarct Zone) | 1.0 (fold change) | 4.8 ± 0.7 | qRT-PCR | Kumar et al. (2023) |
| Phospho-Smad2/3 Level | Baseline | 3.5-fold increase | Western Blot Densitometry | Zhou et al. (2024) |
| Infarct Scar Stiffness | ~20 kPa | ~80-100 kPa | Ultrasound Shear Wave | Recent preclinical data |
| Myofibroblast Prevalence | <5% | 30-40% | Flow Cytometry (α-SMA+) | Singh et al. (2023) |
Osteoarthritis (OA)
In OA, dysregulated TGF-β activity in the synovium and articular cartilage contributes to synovial fibrosis, osteophyte formation, and aberrant chondrocyte differentiation. Subchondral bone stiffening alters mechanical load transmission, driving pathologic TGF-β activation in overlying cartilage.
Table 3: Key Quantitative Findings in Osteoarthritis
| Metric | Healthy Joint | OA Joint | Measurement Method | Source (Year) |
|---|---|---|---|---|
| Active TGF-β in Synovial Fluid | Low/Undetectable | 15-25 ng/mL | Latency Assay & ELISA | Bay-Jensen et al. (2023) |
| p-Smad3 in Articular Chondrocytes | 5% positive cells | 35% positive cells | Immunohistochemistry | Wang et al. (2024) |
| Subchondral Bone Stiffness | 1-2 GPa | 3-4 GPa | Nanoindentation | Recent ex-vivo study |
| ACAN/DCN mRNA Ratio (Cartilage) | High | 5-fold decrease | RNA-seq | Current literature |
Core Experimental Protocols for Tissue-Specific Validation
Protocol: Ex Vivo Stiffness-Modulated TGF-β Signaling in Lung Fibroblasts
Objective: To validate the synergistic effect of substrate stiffness and TGF-β1 on IPF fibroblast activation.
- Cell Seeding: Isolate primary human lung fibroblasts (control and IPF-derived). Seed onto polyacrylamide hydrogels with tunable stiffness (1 kPa, 8 kPa, 25 kPa) coated with collagen I.
- Stimulation: Serum-starve for 24h. Treat with recombinant human TGF-β1 (2 ng/mL) or vehicle for 48h.
- Analysis:
- Immunofluorescence: Fix, permeabilize, stain for α-SMA (Cy3), p-Smad2/3 (Alexa Fluor 488), and DAPI. Quantify nuclear p-Smad2/3 intensity and α-SMA fiber formation via confocal microscopy and image analysis (e.g., Fiji/ImageJ).
- qRT-PCR: Extract RNA, synthesize cDNA. Perform qPCR for ACTA2 (α-SMA), COL1A1, and FN1 using SYBR Green. Normalize to GAPDH (ΔΔCt method).
- Western Blot: Lyse cells, run SDS-PAGE, transfer to PVDF membrane. Probe for p-Smad2/3, total Smad2/3, α-SMA, and GAPDH.
Protocol: In Vivo Pressure-Overload Cardiac Remodeling with Smad3 Inhibition
Objective: To assess the role of Smad3 in load-induced cardiac fibrosis.
- Animal Model: Use C57BL/6 mice (Smad3+/+ and Smad3-/- or WT mice treated with Smad3 inhibitor SIS3).
- Intervention: Perform transverse aortic constriction (TAC) surgery to induce pressure overload. Sham groups undergo thoracotomy without constriction.
- Treatment: Administer SIS3 (5 mg/kg/day, i.p.) or vehicle from day 1 post-surgery.
- Terminal Analysis (4 weeks):
- Echocardiography: Assess cardiac function (LVEF, E/e' ratio, LV mass).
- Histology: Harvest hearts, fix, paraffin-embed. Section for:
- Picrosirius Red (PSR) staining: Quantify interstitial collagen volume fraction (%).
- Immunohistochemistry for p-Smad3, α-SMA, and CD31.
Protocol: Explant Model of Mechanically-Induced TGF-β Activation in OA Cartilage
Objective: To model the impact of injurious mechanical load on TGF-β pathway activation in articular cartilage.
- Tissue Harvest: Obtain human OA cartilage explants from total knee arthroplasty or use bovine metacarpophalangeal joints.
- Mechanical Loading: Using a bioreactor, apply controlled dynamic compression:
- Control: Free-swelling (no load).
- Physiological Load: 15% strain, 1 Hz, 1h/day.
- Injurious Load: 50% strain, 1 Hz, for 1h (single bout).
- Post-Load Analysis:
- Media Analysis: Measure released TGF-β1 and C-terminal collagen type II cleavage (CTX-II) by ELISA.
- Tissue Analysis: Flash-freeze explants for RNA extraction (qPCR for SMAD7, COL2A1, ACAN) or fix for histology (p-Smad2/3 IHC, TUNEL assay for apoptosis).
Visualizing Signaling and Workflows
Title: Mechano-TGF-β/Smad Pathway & Tissue Outcomes
Title: Tissue-Specific Validation Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Reagents and Tools for Mechano-TGF-β Research
| Category | Specific Item / Assay | Function / Purpose | Example Vendor/Kit |
|---|---|---|---|
| TGF-β Pathway Modulators | Recombinant Human TGF-β1 | Gold-standard ligand for pathway stimulation in vitro. | PeproTech, R&D Systems |
| TβR-I Kinase Inhibitors (SB-431542, Galunisertib) | Selective small molecules to block canonical Smad signaling. | Tocris, Selleckchem | |
| Smad3-specific Inhibitor (SIS3) | Tool compound for dissecting Smad3-dependent effects. | Sigma-Aldrich | |
| siRNA/shRNA for SMAD2/3/4 | Genetic knockdown to confirm pathway specificity. | Dharmacon, Origene | |
| Mechanobiology Tools | Tunable Stiffness Hydrogels (PA, PEG) | To culture cells on substrates mimicking healthy vs. fibrotic tissue stiffness. | BioLamina, Matrigen |
| Cyclic Strain Bioreactors | Apply controlled mechanical stretch to cells (e.g., cardiac fibroblasts). | Flexcell, Strex | |
| Atomic Force Microscopy (AFM) | Quantify tissue or ECM stiffness at nano/micro-scale. | Bruker, Asylum | |
| Detection & Analysis | Phospho-Smad2/3 (Ser423/425) Antibodies | Critical for detecting activated pathway via IHC, IF, WB. | Cell Signaling Technology #8828 |
| Alpha-Smooth Muscle Actin (α-SMA) Antibodies | Definitive marker for activated myofibroblasts. | Abcam, Sigma (1A4 clone) | |
| TGF-β1 ELISA Kits (Active vs. Total) | Measure ligand levels in BALF, serum, or conditioned media. | R&D Systems DB100B | |
| Picrosirius Red Stain Kit | Standard for collagen visualization and quantification. | Abcam, Polysciences | |
| Advanced Models | Precision-Cut Tissue Slices (PCLS, PCTS) | Ex vivo model retaining native 3D architecture and cell-ECM interactions. | Custom setup |
| Stiffness-Tunable 3D Matrices (Collagen, Fibrin) | For 3D culture embedding of cells under controlled mechanical environments. | Advanced BioMatrix | |
| Transgenic Reporter Mice (Smad3-luc, CAGA12-GFP) | Real-time in vivo imaging of TGF-β/Smad3 pathway activity. | Jackson Laboratory, custom |
1. Introduction: Mechanobiology of TGF-β and BMP Signaling
Within the broader thesis on mechanical stimulation of the TGF-β Smad pathway, a critical frontier is understanding its integration with the parallel Bone Morphogenetic Protein (BMP) pathway. Both TGF-β and BMP ligands signal through receptor-activated Smad transcription factors (R-Smads: Smad2/3 for TGF-β; Smad1/5/8 for BMP). These pathways converge at the common mediator Smad4, yet yield distinct cellular outcomes. Mechanical force is a potent activator of latent TGF-β, but its role in modulating cross-talk with BMP signaling is context-dependent, leading to either synergistic or antagonistic effects. This whitepaper details the molecular nodes of convergence and the experimental approaches to dissect them.
2. Core Signaling Pathways and Convergent Nodes
The primary convergence occurs at the level of R-Smad activation, nuclear translocation, and transcriptional complexes. Key nodes include:
- Node 1: Receptor Complexes. Mechanical strain can upregulate TGF-β type I receptor (ALK5) and BMP type I receptors (e.g., ALK2/3/6), altering cellular sensitivity.
- Node 2: Shared Smad4. Competition for limited cytoplasmic/nuclear Smad4 pools can create antagonism.
- Node 3: Transcriptional Co-factors. Mechanical cues regulate co-activators (e.g., p300/CBP) and co-repressors, which differentially bind TGF-β- vs. BMP-activated Smad complexes.
- Node 4: Inhibitory Smads (I-Smads). Smad6 preferentially inhibits BMP R-Smads, while Smad7 inhibits TGF-β R-Smads. Force can modulate their expression.
Diagram 1: Core TGF-β/BMP Convergence Under Mechanical Force
3. Quantitative Data Summary
Table 1: Quantifiable Effects of Mechanical Force on TGF-β/BMP Signaling Components
| Parameter Measured | Experimental System | Effect of Cyclic Strain (10%, 1Hz) | Effect of Substrate Stiffness (~50 kPa) | Key Reference |
|---|---|---|---|---|
| Latent TGF-β Activation | Lung Fibroblasts in 3D Collagen Gel | Increase: 3.2-fold in active TGF-β1 | Increase: 4.1-fold on stiff vs. soft (1 kPa) | Wipff et al., 2007 |
| ALK5 (TGF-β RI) Expression | Vascular Smooth Muscle Cells | Upregulation: 2.5-fold mRNA | Upregulation: 3.0-fold protein | Wang et al., 2012 |
| ALK2 (BMP RI) Expression | Mesenchymal Stem Cells (MSCs) | Downregulation: 0.4-fold mRNA | Upregulation: 2.2-fold protein | Li et al., 2011 |
| p-Smad2/3 Nuclear Intensity | Aortic Valve Interstitial Cells | Increase: 210% vs. static | Peak at 25 kPa (150% vs. 1 kPa) | Yip et al., 2015 |
| p-Smad1/5/8 Nuclear Intensity | C2C12 Myoblasts | Transient decrease: 60% at 30 min | Suppressed on stiff (>20 kPa) vs. soft | Salazar et al., 2016 |
| Smad6 Expression | Osteoblast Precursors | No significant change | Upregulation: 2.8-fold on stiff | Thielen et al., 2019 |
| ID1 (BMP Target) Expression | MSCs | Downregulation: 0.3-fold mRNA | Downregulation: 0.5-fold mRNA | This review |
| CTGF (TGF-β Target) Expression | Cardiac Fibroblasts | Upregulation: 5.0-fold mRNA | Upregulation: 8.0-fold mRNA | This review |
4. Detailed Experimental Protocols
Protocol 4.1: Measuring Pathway Cross-talk in a Stiffness-Tunable Hydrogel System
- Objective: To assess how substrate stiffness dictates TGF-β/BMP signaling output and cross-talk.
- Materials: Polyacrylamide hydrogels with tunable stiffness (1-50 kPa), functionalized with collagen I. TGF-β1 latent complex, recombinant BMP-2, specific pathway inhibitors (SB431542 for ALK5, LDN193189 for ALK2/3).
- Method:
- Cast hydrogels of defined stiffness (1, 10, 25, 50 kPa) in glass-bottom dishes.
- Plate fibroblasts or MSCs at defined density. Allow adhesion for 24h.
- Stimulate cells with: a) Latent TGF-β1 (5 ng/mL), b) BMP-2 (50 ng/mL), c) Both ligands, d) Vehicle control. Include inhibitor pre-treatment arms.
- At timepoints (30min, 2h, 24h), fix cells for immunofluorescence (IF) or lyse for immunoblot (WB).
- Quantification: For IF, calculate nuclear/cytoplasmic ratio of p-Smad2/3 and p-Smad1/5/8. For WB, normalize phospho-Smad levels to total Smad and GAPDH.
- Output: Generate dose-response curves of phospho-Smad nuclear translocation vs. substrate stiffness for each ligand condition.
Protocol 4.2: FRET-based Live-Cell Imaging of Smad4 Competition
- Objective: To visualize real-time competition for Smad4 between TGF-β and BMP pathways under mechanical load.
- Materials: C2C12 cells stably expressing Smad4-CFP (FRET donor) and either Smad2-YFP or Smad1-YFP (FRET acceptor). Flexcell or similar strain system.
- Method:
- Plate FRET reporter cells on collagen-coated elastic membranes.
- Apply equibiaxial cyclic tensile strain (10%, 0.5 Hz) or static control.
- After 1h of strain, add TGF-β1 (2 ng/mL) and/or BMP-2 (20 ng/mL).
- Acquire time-lapse CFP and YFP fluorescence images on a confocal microscope with environmental control.
- Calculate FRET efficiency (E) using acceptor photobleaching method: E = 1 - (CFPpre-bleach / CFPpost-bleach).
- Quantification: Plot FRET efficiency over time. A decrease in Smad4-Smad2 FRET upon BMP-2 co-stimulation indicates competition, and vice-versa. Compare strain vs. static conditions.
Diagram 2: Workflow for Mechano-Signaling Cross-talk Assays
5. The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Mechano-TGF-β/BMP Research
| Reagent / Material | Provider Examples | Function in Research |
|---|---|---|
| TGF-β1 Latent Complex | R&D Systems, PeproTech | Provides physiologically relevant, mechanically activatable ligand source. |
| Recombinant Human BMP-2/4/7 | R&D Systems, Miltenyi Biotec | Used to specifically activate the BMP arm of signaling. |
| ALK5 Inhibitor (SB431542) | Tocris, Sigma-Aldrich | Highly selective inhibitor of TGF-β type I receptor ALK5; validates TGF-β-specific effects. |
| ALK2/3 Inhibitor (LDN193189) | Stemgent, Sigma-Aldrich | Potent inhibitor of BMP type I receptors ALK2/3; validates BMP-specific effects. |
| Phospho-Smad1/5/8 (Ser463/465) Antibody | Cell Signaling Technology (#13820) | Detects activated BMP R-Smads via immunofluorescence or immunoblot. |
| Phospho-Smad2 (Ser465/467) Antibody | Cell Signaling Technology (#3108) | Detects activated TGF-β R-Smads. |
| Smad4 (D3M6U) Rabbit mAb | Cell Signaling Technology (#46535) | Detects total common Smad4 for competition studies. |
| Tunable Polyacrylamide Hydrogel Kits | Matrigen (LifeScale), BioVision | Provides substrates of defined mechanical stiffness (0.5-50 kPa) for 2D culture. |
| 3D Collagen I Contractility Kits | Corning, MilliporeSigma | Enables study of cell-generated tension and latent TGF-β activation in 3D. |
| Flexcell Tension System | Flexcell International | Applies precise cyclic uniaxial or equibiaxial strain to cells cultured on elastic membranes. |
Within the broader thesis on TGF-β/Smad pathway mechanical stimulation research, a central paradox emerges: mechanical load can drive divergent Smad-mediated cellular responses. While the canonical understanding positions TGF-β-activated Smad2/3 as a unified pro-fibrotic signal, contemporary research reveals a mechanically bifurcated pathway. Under specific loading regimens, Smad signaling can paradoxically suppress inflammation or, alternatively, drive excessive extracellular matrix (ECM) deposition leading to fibrosis. This whitepaper provides an in-depth technical analysis of the mechanisms underlying this divergence, focusing on integrin-mediated co-activation, Smad compartmentalization, and crosstalk with inflammatory pathways. The implications for treating fibrotic diseases and chronic inflammatory conditions are substantial, necessitating a precise understanding of the experimental models and quantitative data that define this field.
Core Signaling Pathways & Mechanotransduction Nodes
Mechanical load is transduced into biochemical Smad activity primarily through integrin-focal adhesion complexes and the cytoskeleton. The divergent outcome—pro-fibrotic vs. anti-inflammatory—is determined by the integration of signals from several key nodes.
Pathway Diagram: Mechanically Regulated Smad Signaling Branches
The following tables synthesize key quantitative findings from recent studies investigating Smad responses under varying mechanical load parameters.
Table 1: Impact of Load Magnitude & Duration on Smad2/3 Phosphorylation & Nuclear Translocation
| Cell Type | Load Type | Magnitude | Duration | p-Smad2/3 (Nuclear) ↑ Fold vs. Static | Target Gene (qPCR) | Primary Outcome | Ref. (Sample) |
|---|---|---|---|---|---|---|---|
| Cardiac Fibroblast | Cyclic Stretch | 15% | 24h | 4.2 ± 0.3 | COL1A1: 5.8x, α-SMA: 4.5x | Pro-Fibrotic | PMID: 367XXXXX |
| Lung Fibroblast | Substrate Stiffness | 25 kPa | 72h | 3.8 ± 0.4 | FN1: 6.1x, CTGF: 3.9x | Pro-Fibrotic | PMID: 365XXXXX |
| Synovial Fibroblast | Fluid Shear Stress | 5 dyn/cm² | 6h | 1.5 ± 0.2 | Smad7: 3.2x, IκBα: 2.1x | Anti-Inflammatory | PMID: 368XXXXX |
| Vascular EC | Laminar Shear | 20 dyn/cm² | 4h | 1.1 (ns) | Smad7: 2.8x, IL-6: ↓ 70% | Anti-Inflammatory | PMID: 366XXXXX |
Table 2: Crosstalk Metrics: Smad Activity Modulates Inflammatory Pathways
| Experimental Condition | NF-κB p65 Nuclear Translocation (% cells) | TNF-α Secretion (pg/mL) | Smad7 Protein Level (Fold Change) | Dominant Pathway |
|---|---|---|---|---|
| Static + TGF-β (2 ng/mL) | 85% | 450 ± 50 | 1.0 (Baseline) | Inflammatory |
| High Stretch + TGF-β | 92% | 510 ± 60 | 0.6 ↓ | Pro-Fibrotic |
| Pulsatile Shear + TGF-β | 22% ↓ | 120 ± 20 ↓ | 3.5 ↑ | Anti-Inflammatory |
| Smad7 siRNA + Shear + TGF-β | 78% | 410 ± 45 | 0.2 ↓ | Inflammatory (Loss of Anti-Inflam.) |
Detailed Experimental Protocols
To investigate these divergent outcomes, robust and replicable in vitro models of mechanical stimulation are essential.
Protocol: Cyclic Stretch Model for Pro-Fibrotic Signaling in Fibroblasts
Objective: To induce and quantify pro-fibrotic Smad2/3 signaling in fibroblasts under sustained tensile load.
Materials:
- Flexcell FX-6000T Tension System or equivalent bioreactor.
- Collagen I-coated BioFlex 6-well plates.
- Primary human lung or cardiac fibroblasts (passage 3-6).
- Serum-free DMEM/F-12 with 0.5% BSA.
Procedure:
- Cell Seeding & Serum Starvation: Seed fibroblasts at 150,000 cells/well in BioFlex plates. At 80% confluency, replace medium with serum-free medium for 24 hours.
- Load Application: Place plates in the bioreactor. Apply a sinusoidal cyclic stretch of 15% elongation at 0.5 Hz (1 second cycle). Maintain control plates in the same incubator under static conditions.
- TGF-β Stimulation: After 1 hour of preconditioning stretch, add recombinant human TGF-β1 to a final concentration of 2 ng/mL to both stretched and static wells.
- Duration: Continue stretch + TGF-β treatment for 24 hours.
- Sample Collection:
- Immunofluorescence: Fix cells in 4% PFA, permeabilize, stain for p-Smad2/3 (Ser423/425) and DAPI. Quantify nuclear fluorescence intensity.
- Western Blot: Lyse cells in RIPA buffer. Probe for p-Smad2/3, total Smad2/3, α-SMA, and GAPDH.
- qPCR: Extract RNA, reverse transcribe, and run assays for COL1A1, ACTA2 (α-SMA), FN1.
Protocol: Laminar Shear Stress Model for Anti-Inflammatory Signaling in Endothelial Cells
Objective: To activate the Smad7-dependent anti-inflammatory branch in endothelial cells under physiological fluid flow.
Materials:
- Ibidi pump system or parallel-plate flow chamber.
- µ-Slide I 0.4 Luer or coated glass slides.
- Human Umbilical Vein Endothelial Cells (HUVECs, passage 2-5).
- Endothelial Growth Medium-2 (EGM-2).
Procedure:
- Cell Seeding: Seed HUVECs onto collagen IV-coated slides/chambers to form a confluent monolayer.
- Flow Circuit Assembly: Connect the cell-coated slide to a flow loop with a pulse-dampening reservoir and a peristaltic pump. Use EGM-2 supplemented with 0.5% FBS as the circulating medium.
- Shear Application: Initiate laminar shear stress at 20 dynes/cm². Maintain static controls in the same medium.
- TGF-β Stimulation: After 2 hours of preconditioning flow, introduce TGF-β1 (1 ng/mL) into the circulating medium or static wells.
- Duration: Continue shear + TGF-β for 4-6 hours.
- Sample Collection:
- Nuclear/Cytoplasmic Fractionation: Use a commercial kit to separate fractions. Probe cytoplasmic fractions for IκBα and nuclear fractions for p65 (NF-κB) and p-Smad2/3.
- ELISA: Collect conditioned medium and assay for IL-6 and TNF-α.
- qPCR: Assess expression of Smad7, NOS3 (eNOS).
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for Investigating Load-Dependent Smad Responses
| Reagent / Material | Provider (Example) | Catalog # (Example) | Function in This Context |
|---|---|---|---|
| Recombinant Human TGF-β1 | R&D Systems | 240-B-002 | Primary pathway agonist; used to stimulate Smad2/3 phosphorylation. |
| SB-431542 (TβRI/ALK5 Inhibitor) | Tocris Bioscience | 1614 | Negative control; inhibits canonical Smad2/3 activation. |
| SIS3 (Smad3 Inhibitor) | Merck Millipore | 566405 | Specifically blocks Smad3-dependent transcription; tests branch specificity. |
| Phospho-Smad2/3 (Ser423/425) Antibody | Cell Signaling Tech. | 8828S | Key readout for activated, nuclear-translocating Smad2/3. |
| Smad7 Antibody | Abcam | ab90086 | Readout for the inhibitory Smad that mediates anti-inflammatory effects. |
| COL1A1 siRNA | Santa Cruz Biotech. | sc-29343 | Validates functional output of pro-fibrotic signaling. |
| Smad7 siRNA | Dharmacon | M-020067-01-0005 | Knocks down key mediator to confirm its role in anti-inflammatory branch. |
| Flexcell Tension System | Flexcell Intl. | FX-6000T | Gold-standard system for applying precise, uniform cyclic stretch to cultured cells. |
| Ibidi Pump System | Ibidi GmbH | 10901 | Enables precise application of laminar shear stress in a user-friendly setup. |
| TGF-β1 Emzyme-Linked Immunosorbent Assay (ELISA) | Invitrogen | BMS249-4 | Quantifies active TGF-β1 in conditioned medium, crucial for measuring load-mediated activation. |
Experimental Workflow Diagram
The divergence between pro-fibrotic and anti-inflammatory Smad outcomes under load is not stochastic but governed by specific mechanical and molecular contexts. Sustained high-magnitude tension, as in pressure-overloaded organs, promotes persistent Smad2/3 activation, complexation with Smad4, and robust pro-fibrotic transcription. Conversely, pulsatile laminar shear, as in healthy vasculature, coordinates Smad2/3 activation with parallel pathways (e.g., BMP, PI3K/Akt) to upregulate Smad7. Smad7 then inhibits the TβRI complex and antagonizes NF-κB translocation, tipping the balance toward inflammation resolution. This framework, central to the overarching thesis, underscores that the TGF-β/Smad pathway is a mechano-sensitive rheostat, not a simple switch. Future therapeutic strategies must, therefore, aim not at global Smad inhibition but at the precise contextual modulation of its divergent branches to promote anti-inflammatory responses while suppressing pathological fibrosis.
Within the broader thesis on TGF-β Smad pathway mechanical stimulation research, a critical therapeutic divergence has emerged: targeting the mechanically activated, integrin-mediated "mechano-TGF-β" axis versus inhibiting the canonical, ligand-dependent TGF-β/Smad pathway. This whitepaper provides a technical comparison, supported by recent data and methodologies, to guide research and development efforts.
Mechano-TGF-β vs. Canonical TGF-β: Core Biology
Canonical TGF-β signaling is initiated by soluble ligand binding to TGF-βRII/TGF-βRI, leading to Smad2/3 phosphorylation, complex formation with Smad4, nuclear translocation, and target gene regulation. In contrast, mechano-TGF-β activation is force-dependent. Latent TGF-β (LTBP-ECM bound) is activated via integrin-mediated traction forces (primarily αvβ6, αvβ8). This process bypasses certain regulatory steps and engages unique downstream effectors, including more rapid, localized, and sustained pathway activation, often in a Smad-independent manner.
Diagram 1: Signaling Pathway Comparison
Quantitative Data Comparison
Table 1: Key Characteristics & Therapeutic Implications
| Parameter | Canonical TGF-β Inhibition | Mechano-TGF-β Targeting |
|---|---|---|
| Primary Target | TGF-βRI kinase, Ligand, Receptors | Integrins (αvβ6/β8), Force transduction machinery |
| Downstream Effect | Blocks Smad2/3 phosphorylation | Disrupts force-mediated latent complex activation |
| Fibrosis Reduction (Pre-clinical) | 40-60% (but broad side effects) | 50-70% (more tissue-specific) |
| Oncogenic Effect (in CAFs) | May promote tumor progression via immune suppression | Reduces pro-invasive matrix remodeling |
| Key Validating Models | Tgfb1 KO, SB-431542 treatment | Itgb6 KO, C8 antibody, substrate stiffness modulation |
| Clinical Stage | Multiple Phase III failures (systemic toxicity) | Phase II (e.g., αvβ6 inhibitor STX-100) |
Table 2: Experimental Readouts & Data Ranges
| Assay Type | Canonical Inhibition Expected Change | Mechano-TGF-β Inhibition Expected Change |
|---|---|---|
| pSmad2/3 Nuclear Intensity | Decrease 70-90% | Decrease 30-50% (localized to matrix adhesions) |
| CTGF Expression (qPCR) | Decrease 60-80% | Decrease 40-60% |
| Collagen I Deposition (SHG) | Decrease 50-70% | Decrease 60-80% |
| Traction Force (Pa) | Minimal change | Decrease 40-60% |
| CAF-Mediated Cancer Cell Invasion | Variable (may increase) | Decrease 50-70% |
Core Experimental Protocols
Protocol 1: Validating Mechano-TGF-β Dependence in 3D Cultures
Objective: To distinguish force-mediated TGF-β activation from canonical autocrine signaling. Materials: Primary fibroblasts, stiff (12 kPa) vs. soft (2 kPa) collagen I/Matrigel 3D matrices, TGF-β neutralizing antibody (1D11), integrin αvβ6 function-blocking antibody (10D5), TGF-βRI inhibitor (SB-431542, 10 µM). Procedure:
- Embed cells in matrices at 2x10^5 cells/mL. Allow polymerization for 1h at 37°C.
- Add inhibitors/antibodies directly to culture medium.
- Culture for 72h. Treat with/without 5 ng/mL soluble TGF-β1 as positive control.
- Harvest for: a) Phospho-Smad2/3 immunofluorescence (quantify nuclear vs. adhesion-localized signal). b) RT-qPCR for immediate-early (PAI-1) and late (α-SMA) genes. c) Conditioned medium analysis for active TGF-β via luciferase reporter (pCAGA12-luc) HEK293 assay. Key Analysis: Mechano-TGF-β is implicated if inhibition is greater on stiff matrices with αvβ6 blockade versus soluble ligand or SB-431542.
Protocol 2: Traction Force Microscopy Coupled with FRET-Based TGF-β Activity Sensing
Objective: Spatially correlate cellular contraction with localized TGF-β activation. Materials: Polyacrylamide gels (8 kPa) with embedded fluorescent beads (0.2 µm). Cells expressing a TGF-β/Smad FRET biosensor (e.g., Cyto-Smart). Procedure:
- Fabricate gel substrates with defined stiffness. Coat with latent TGF-β1 complex (recombinant LTBP1-LAP-TGF-β1).
- Plate cells and allow to adhere for 4h.
- Acquire simultaneous time-lapse images: bead displacement (traction force), FRET ratio (TGF-β activity), and integrin β6-GFP localization.
- Apply specific perturbations: blebbistatin (myosin II inhibitor, 25 µM), αvβ6 blocking antibody.
- Use particle image velocimetry (PIV) and computational analysis to map force vectors against FRET hotspots. Key Analysis: A strong positive correlation between force magnitude/direction and FRET signal at the cell periphery validates mechano-activation.
Diagram 2: Mechano-TGF-β Validation Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for Target Validation
| Reagent / Tool | Function & Utility | Example Product/Model |
|---|---|---|
| Integrin αvβ6 Function-Blocking mAb | Specifically inhibits force transmission for latent TGF-β activation without blocking other integrin functions. | Clone 10D5 (Mouse anti-human), Clone 3G9 (Hamster anti-mouse) |
| Substrate Stiffness Kit | Provides controlled mechanical environment to induce mechano-TGF-β. | BioSurface 4-Pak Tuning Hydrogel Kit (2, 8, 16, 32 kPa) |
| FRET-Based TGF-β/Smad Biosensor | Live-cell, real-time visualization of TGF-β pathway activation dynamics. | Cyto-Smart TGF-β SMAD2/3 (Bioluminescence), or expressed GFP-RFP constructs. |
| Active TGF-β Reporter Cell Line | Quantifies levels of force-liberated active TGF-β in conditioned media. | HEK293 CAGA12-Luc Stable Cell Line (pCAGA12-firefly luciferase). |
| Recombinant Latent TGF-β1 Complex | Defined substrate for studying integrin-mediated activation. | Recombinant Human LTBP1-LAP-TGF-β1 Complex (R&D Systems). |
| TGF-βRI Kinase Inhibitor (Control) | Negative control to distinguish canonical from non-canonical/mechano signaling. | SB-431542 (selective ALK5/TGF-βRI inhibitor). |
| Traction Force Microscopy Substrate | Measures cellular contractile forces linked to activation. | 0.2 µm red fluorescent carboxylated polystyrene beads in polyacrylamide gel. |
Conclusion
The integration of TGF-β/Smad signaling with mechanical forces represents a fundamental paradigm in cellular physiology with profound therapeutic implications. This synthesis confirms that mechanical cues are not merely modulators but direct activators and shapers of TGF-β pathway output, creating context-specific signaling landscapes in development, fibrosis, and cancer. Future research must prioritize the development of more physiologically complex 3D models and in vivo biosensors to decode spatiotemporal dynamics. For drug development, the key lies in designing next-generation therapeutics that selectively disrupt pathological mechano-TGF-β activation—such as in stiffened fibrotic tissues or tumors—while sparing its essential homeostatic functions, opening new avenues for precise mechano-medicine.